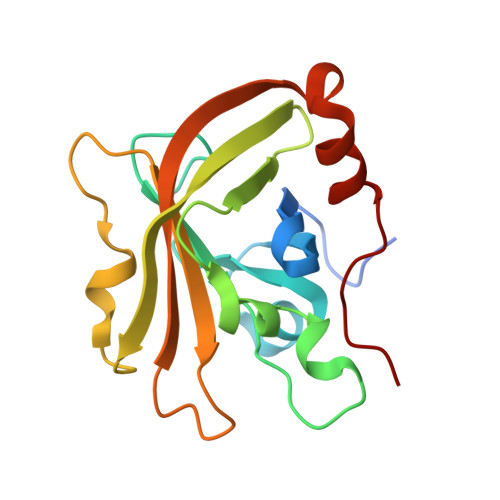
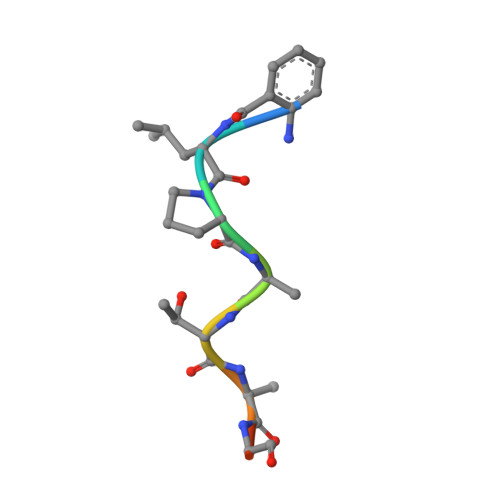
Structures of Streptococcus pyogenes class A sortase in complex with substrate and product mimics provide key details of target recognition.
Johnson, D.A., Piper, I.M., Vogel, B.A., Jackson, S.N., Svendsen, J.E., Kodama, H.M., Lee, D.E., Lindblom, K.M., McCarty, J., Antos, J.M., Amacher, J.F.(2022) J Biol Chem 298: 102446-102446
- PubMed: 36055407
- DOI: https://doi.org/10.1016/j.jbc.2022.102446
- Primary Citation of Related Structures:
7S4O, 7S51, 7T8Y, 7T8Z - PubMed Abstract:
The cell wall is a critical extracellular barrier for bacteria and many other organisms. In bacteria, this structural layer consists of peptidoglycan, which maintains cell shape and structural integrity and provides a scaffold for displaying various protein factors. To attach proteins to the cell wall, Gram-positive bacteria utilize sortase enzymes, which are cysteine transpeptidases that recognize and cleave a specific sorting signal, followed by ligation of the sorting signal-containing protein to the peptidoglycan precursor lipid II (LII). This mechanism is the subject of considerable interest as a target for therapeutic intervention and as a tool for protein engineering, where sortases have enabled sortase-mediated ligation or sortagging strategies. Despite these uses, there remains an incomplete understanding of the stereochemistry of substrate recognition and ligation product formation. Here, we solved the first structures of sortase A from Streptococcus pyogenes bound to two substrate sequences, LPATA and LPATS. In addition, we synthesized a mimetic of the product of sortase-mediated ligation involving LII (LPAT-LII) and solved the complex structure in two ligand conformations. These structures were further used as the basis for molecular dynamics simulations to probe sortase A-ligand dynamics and to construct a model of the acyl-enzyme intermediate, thus providing a structural view of multiple key states in the catalytic mechanism. Overall, this structural information provides new insights into the recognition of the sortase substrate motif and LII ligation partner and will support the continued development of sortases for protein engineering applications.
Organizational Affiliation:
Department of Chemistry, Western Washington University, Bellingham, Washington, USA.