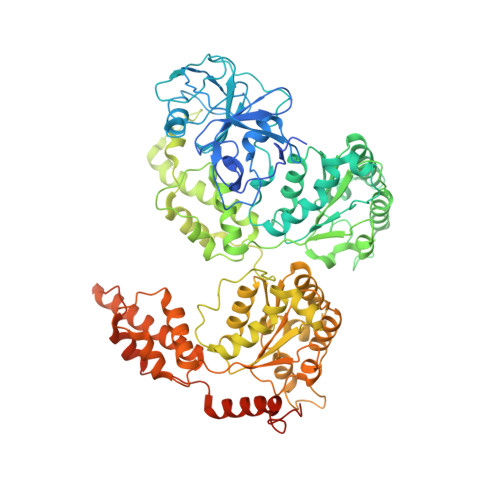
AAA+ ATPase p97/VCP mutants and inhibitor binding disrupt inter-domain coupling and subsequent allosteric activation.
Caffrey, B., Zhu, X., Berezuk, A., Tuttle, K., Chittori, S., Subramaniam, S.(2021) J Biol Chem 297: 101187-101187
- PubMed: 34520757
- DOI: https://doi.org/10.1016/j.jbc.2021.101187
- Primary Citation of Related Structures:
7RL6, 7RL7, 7RL9, 7RLA, 7RLB, 7RLC, 7RLD, 7RLF, 7RLG, 7RLH, 7RLI, 7RLJ - PubMed Abstract:
The human AAA+ ATPase p97, also known as valosin-containing protein, a potential target for cancer therapeutics, plays a vital role in the clearing of misfolded proteins. p97 dysfunction is also known to play a crucial role in several neurodegenerative disorders, such as MultiSystem Proteinopathy 1 (MSP-1) and Familial Amyotrophic Lateral Sclerosis (ALS). However, the structural basis of its role in such diseases remains elusive. Here, we present cryo-EM structural analyses of four disease mutants p97 R155H , p97 R191Q , p97 A232E , p97 D592N , as well as p97 E470D , implicated in resistance to the drug CB-5083, a potent p97 inhibitor. Our cryo-EM structures demonstrate that these mutations affect nucleotide-driven allosteric activation across the three principal p97 domains (N, D1, and D2) by predominantly interfering with either (1) the coupling between the D1 and N-terminal domains (p97 R155H and p97 R191Q ), (2) the interprotomer interactions (p97 A232E ), or (3) the coupling between D1 and D2 nucleotide domains (p97 D592N , p97 E470D ). We also show that binding of the competitive inhibitor, CB-5083, to the D2 domain prevents conformational changes similar to those seen for mutations that affect coupling between the D1 and D2 domains. Our studies enable tracing of the path of allosteric activation across p97 and establish a common mechanistic link between active site inhibition and defects in allosteric activation by disease-causing mutations and have potential implications for the design of novel allosteric compounds that can modulate p97 function.
Organizational Affiliation:
Department of Biochemistry and Molecular Biology, University of British Columbia, Vancouver, British Columbia, Canada.