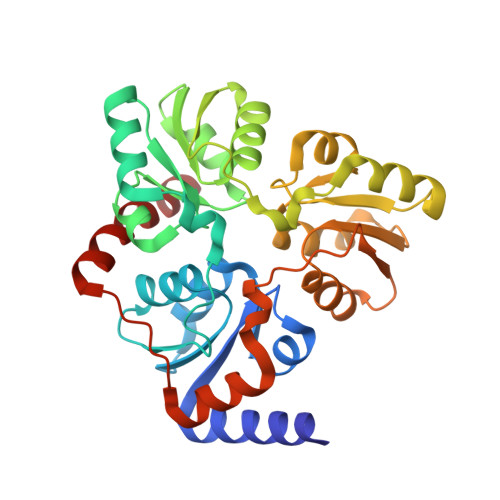
Transient Formation of a Second Active Site Cavity during Quinolinic Acid Synthesis by NadA.
Basbous, H., Volbeda, A., Amara, P., Rohac, R., Martin, L., Ollagnier de Choudens, S., Fontecilla-Camps, J.C.(2021) ACS Chem Biol 16: 2423-2433
- PubMed: 34609124
- DOI: https://doi.org/10.1021/acschembio.1c00541
- Primary Citation of Related Structures:
7P4M, 7P4P, 7P4Q - PubMed Abstract:
Quinolinate synthase, also called NadA, is a [4Fe-4S]-containing enzyme that uses what is probably the oldest pathway to generate quinolinic acid (QA), the universal precursor of the biologically essential cofactor nicotinamide adenine dinucleotide (NAD). Its synthesis comprises the condensation of dihydroxyacetone phosphate (DHAP) and iminoaspartate (IA), which involves dephosphorylation, isomerization, cyclization, and two dehydration steps. The convergence of the three homologous domains of NadA defines a narrow active site that contains a catalytically essential [4Fe-4S] cluster. A tunnel, which can be opened or closed depending on the nature (or absence) of the bound ligand, connects this cofactor to the protein surface. One outstanding riddle has been the observation that the so far characterized active site is too small to bind IA and DHAP simultaneously. Here, we have used site-directed mutagenesis, X-ray crystallography, functional analyses, and molecular dynamics simulations to propose a condensation mechanism that involves the transient formation of a second active site cavity to which one of the substrates can migrate before this reaction takes place.
Organizational Affiliation:
Univ. Grenoble Alpes, CNRS, CEA, Laboratoire de Chimie et Biologie des Métaux, BioCat, 38000 Grenoble, France.