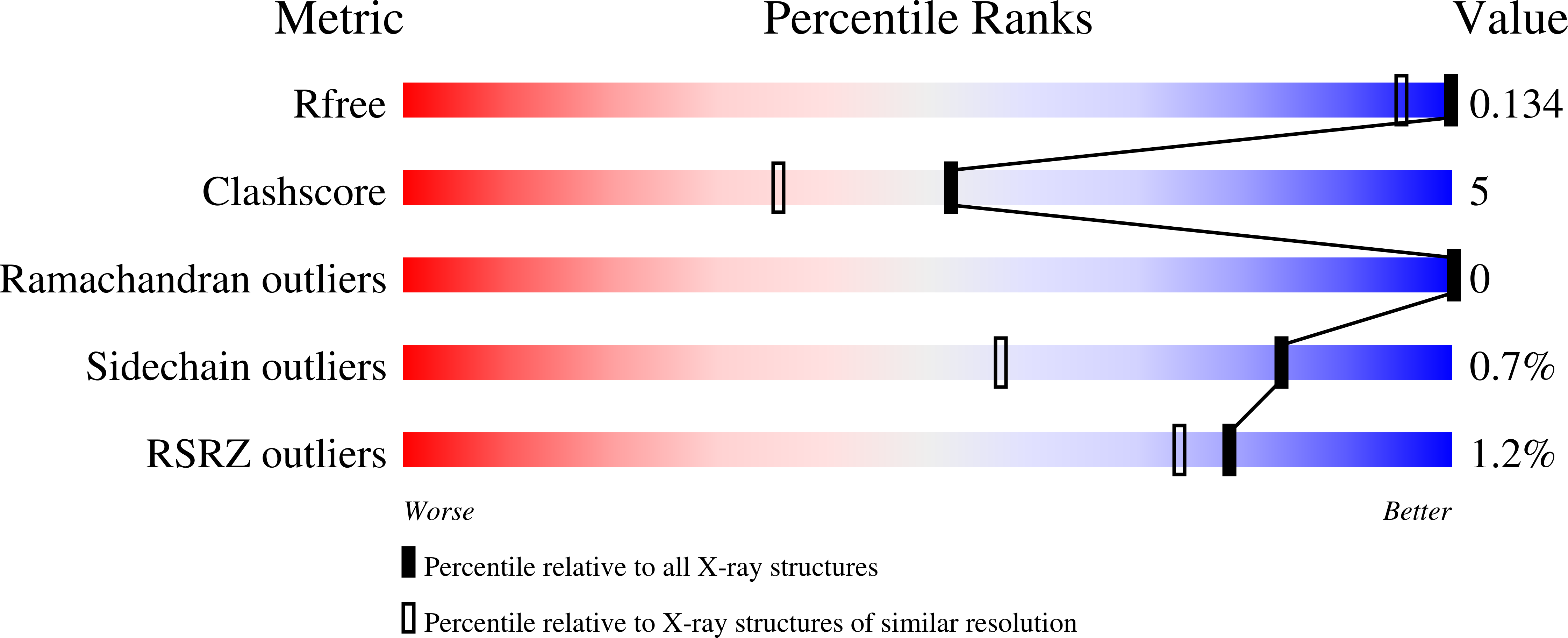
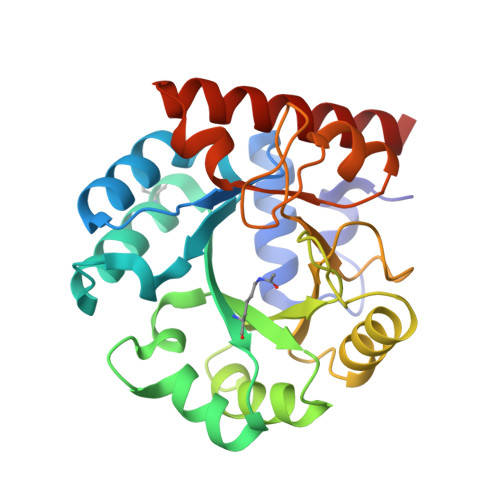
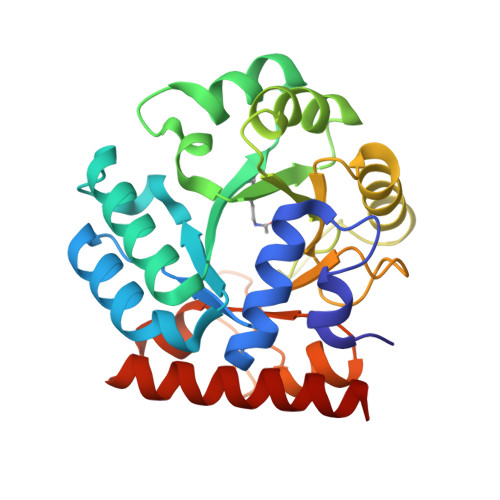
Ground-state destabilization by electrostatic repulsion is not a driving force in orotidine-5-monophosphate decarboxylase catalysis
Rindfleisch, S., Krull, M., Uranga, J., Schmidt, T., Rabe von Pappenheim, F., Kirck, L.L., Balouri, A., Schneider, T., Chari, A., Kluger, R., Bourenkov, G., Diederichsen, U., Mata, R.A., Tittmann, K.(2022) Nat Catal 5: 332-341