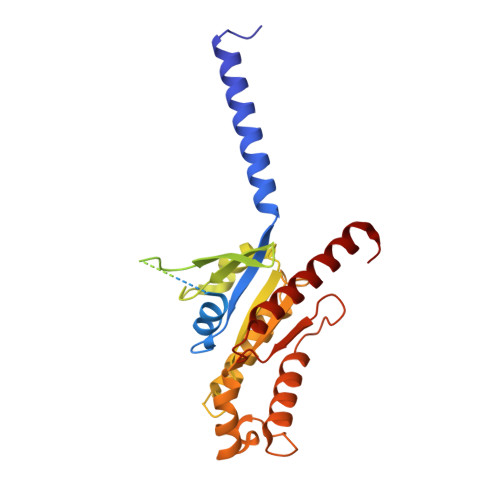
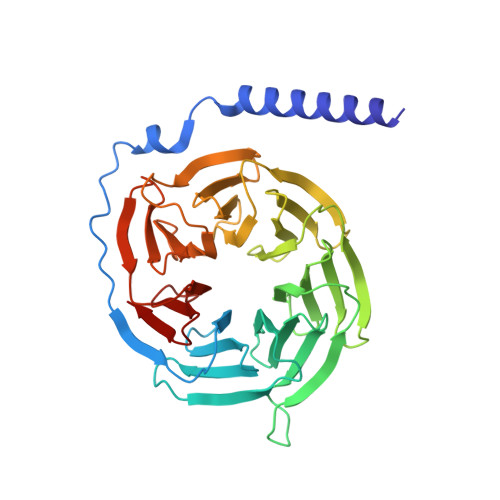
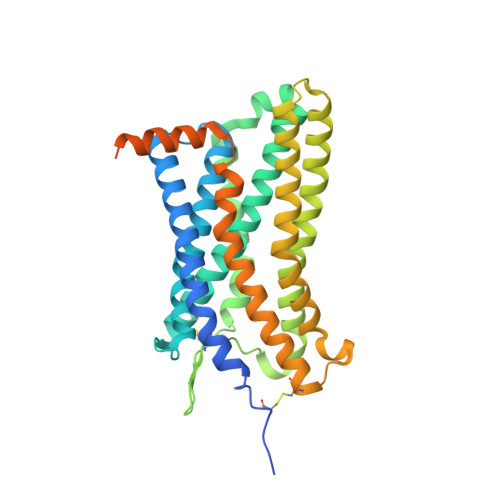
Structural basis of the activation of the CC chemokine receptor 5 by a chemokine agonist.
Isaikina, P., Tsai, C.J., Dietz, N., Pamula, F., Grahl, A., Goldie, K.N., Guixa-Gonzalez, R., Branco, C., Paolini-Bertrand, M., Calo, N., Cerini, F., Schertler, G.F.X., Hartley, O., Stahlberg, H., Maier, T., Deupi, X., Grzesiek, S.(2021) Sci Adv 7
- PubMed: 34134983
- DOI: https://doi.org/10.1126/sciadv.abg8685
- Primary Citation of Related Structures:
7O7F - PubMed Abstract:
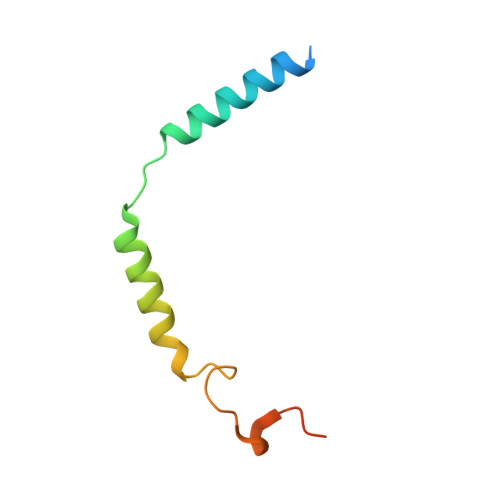
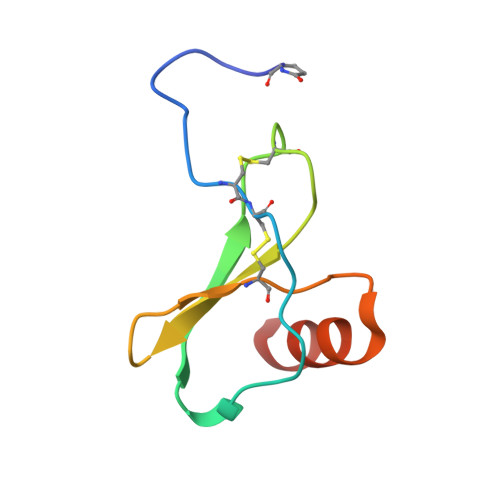
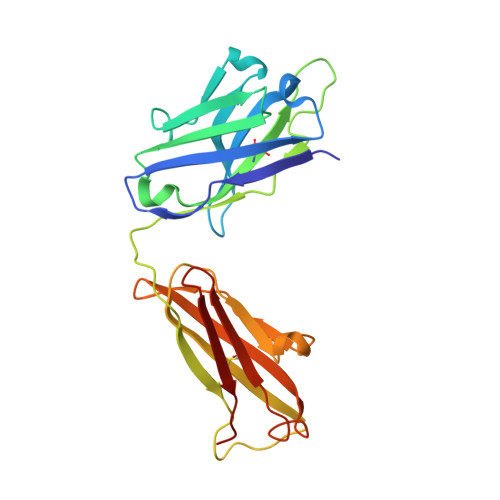
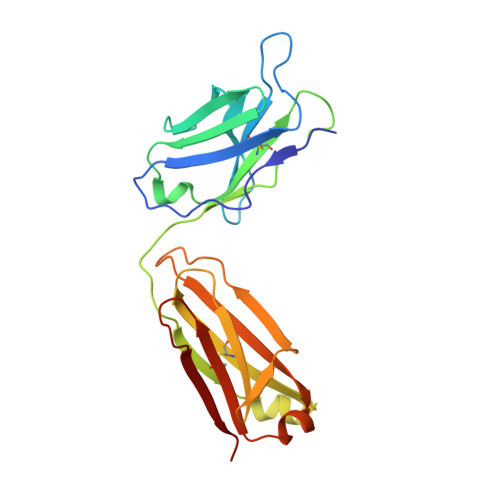
The human CC chemokine receptor 5 (CCR5) is a G protein-coupled receptor (GPCR) that plays a major role in inflammation and is involved in cancer, HIV, and COVID-19. Despite its importance as a drug target, the molecular activation mechanism of CCR5, i.e., how chemokine agonists transduce the activation signal through the receptor, is yet unknown. Here, we report the cryo-EM structure of wild-type CCR5 in an active conformation bound to the chemokine super-agonist [6P4]CCL5 and the heterotrimeric G i protein. The structure provides the rationale for the sequence-activity relation of agonist and antagonist chemokines. The N terminus of agonist chemokines pushes onto specific structural motifs at the bottom of the orthosteric pocket that activate the canonical GPCR microswitch network. This activation mechanism differs substantially from other CC chemokine receptors that bind chemokines with shorter N termini in a shallow binding mode involving unique sequence signatures and a specialized activation mechanism.
Organizational Affiliation:
Focal Area Structural Biology and Biophysics, Biozentrum, University of Basel, CH-4056 Basel, Switzerland.