Overcoming the Pregnane X Receptor Liability: Rational Design to Eliminate PXR-Mediated CYP Induction.
Ramanjulu, J.M., Williams, S.P., Lakdawala, A.S., DeMartino, M.P., Lan, Y., Marquis, R.W.(2021) ACS Med Chem Lett 12: 1396-1404
- PubMed: 34531948
- DOI: https://doi.org/10.1021/acsmedchemlett.1c00187
- Primary Citation of Related Structures:
7N2A, 7RIO, 7RIU, 7RIV - PubMed Abstract:
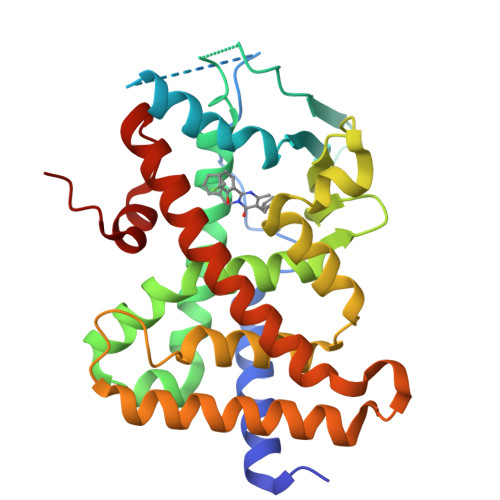
The pregnane X receptor (PXR) regulates expression of proteins responsible for all three phases required for the detoxification mechanism, which include CYP450 enzymes, phase II enzymes, and multidrug efflux pumps. Therefore, PXR is a prominent receptor that is responsible for xenobiotic excretion and drug-drug interactions. Pyrimidinone 1 is an antagonist of the calcium sensing receptor (CaSR) and a strong activator of PXR. Repeat oral administration revealed diminished exposures over time, which prohibited further progression. A medicinal chemistry campaign was initiated to understand and abolish activation of PXR in order to increase systemic exposures. Rational structure-activity relationship investigations utilizing cocrystal structures and a de novo pharmacophore model resulted in compounds devoid of PXR activation. These studies culminated in the first orally active CaSR antagonist 8 suitable for progression. Cocrystallography, the pharmacophore model employed, and additional observations reported herein supported rational elimination of PXR activation and have applicability across diverse chemical classes to help erase PXR-driven drug-drug interactions.
Organizational Affiliation:
GlaxoSmithKline, 1250 South Collegeville Road, Collegeville, Pennsylvania 19426, United States.