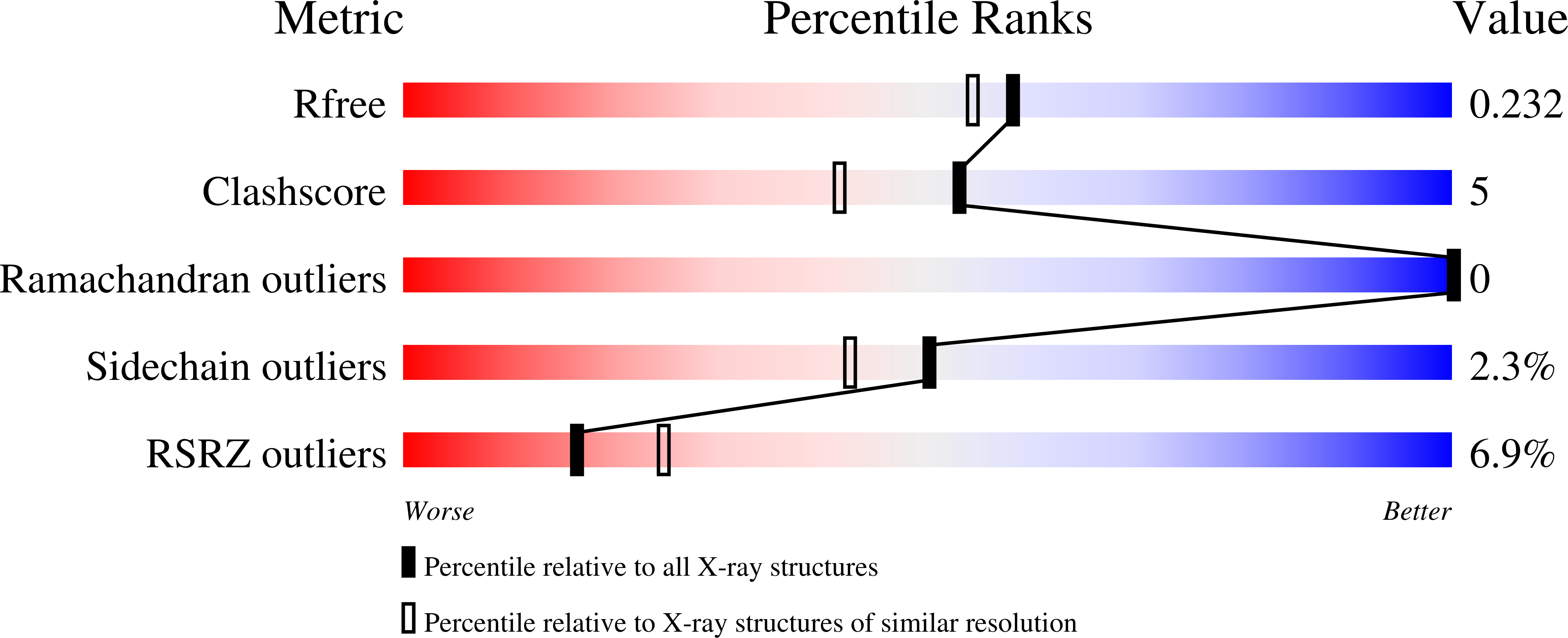
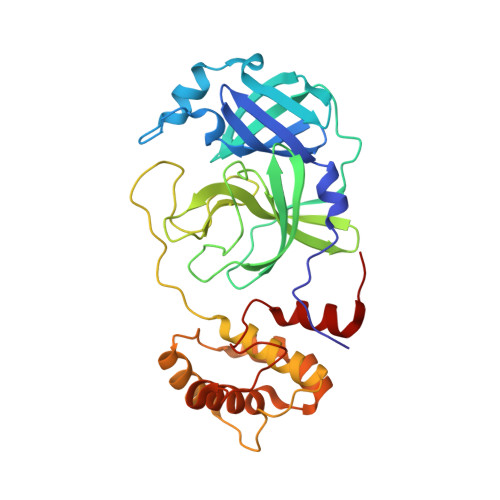
Structure-guided design of a perampanel-derived pharmacophore targeting the SARS-CoV-2 main protease.
Deshmukh, M.G., Ippolito, J.A., Zhang, C.H., Stone, E.A., Reilly, R.A., Miller, S.J., Jorgensen, W.L., Anderson, K.S.(2021) Structure 29: 823
- PubMed: 34161756
- DOI: https://doi.org/10.1016/j.str.2021.06.002
- Primary Citation of Related Structures:
7M8M, 7M8N, 7M8O, 7M8P, 7M8X, 7M8Y, 7M8Z, 7M90, 7M91 - PubMed Abstract:
There is a clinical need for direct-acting antivirals targeting SARS-CoV-2, the coronavirus responsible for the COVID-19 pandemic, to complement current therapeutic strategies. The main protease (M pro ) is an attractive target for antiviral therapy. However, the vast majority of protease inhibitors described thus far are peptidomimetic and bind to the active-site cysteine via a covalent adduct, which is generally pharmacokinetically unfavorable. We have reported the optimization of an existing FDA-approved chemical scaffold, perampanel, to bind to and inhibit M pro noncovalently with IC 50 s in the low-nanomolar range and EC 50 s in the low-micromolar range. Here, we present nine crystal structures of M pro bound to a series of perampanel analogs, providing detailed structural insights into their mechanism of action and structure-activity relationship. These insights further reveal strategies for pursuing rational inhibitor design efforts in the context of considerable active-site flexibility and potential resistance mechanisms.
Organizational Affiliation:
Medical Scientist Training Program (MD-PhD), Yale School of Medicine, New Haven, CT, USA; Department of Pharmacology, Yale University School of Medicine, New Haven, CT 06520-8066, USA.