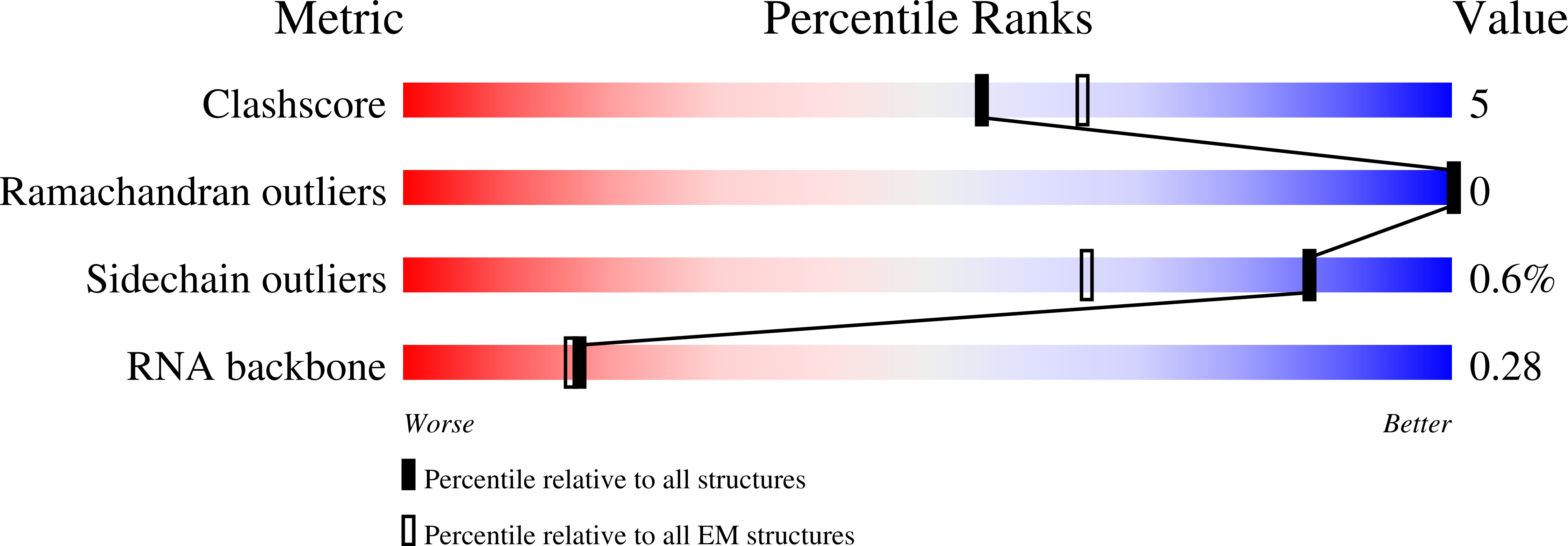
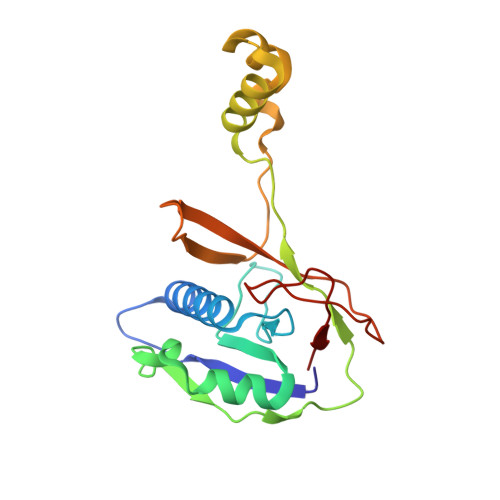
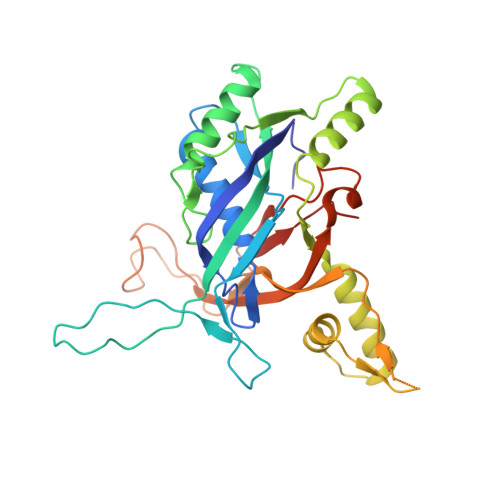
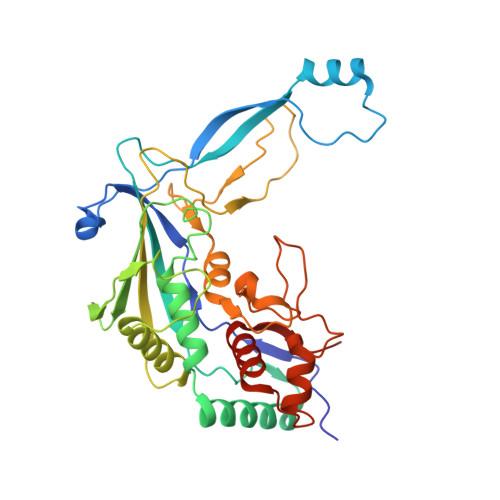
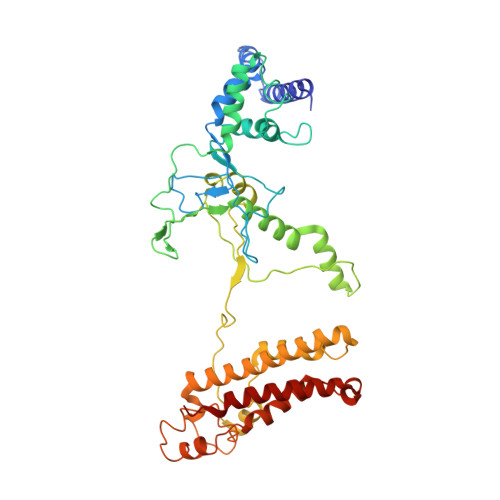
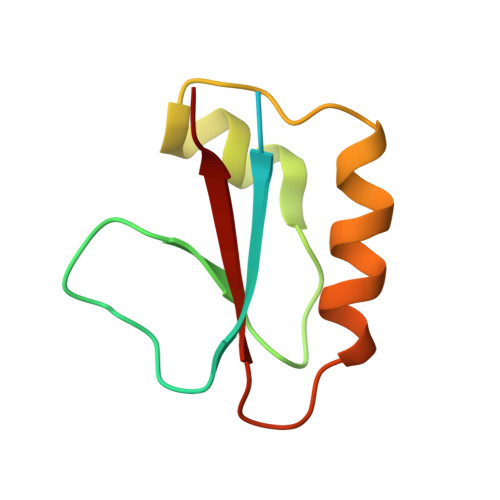
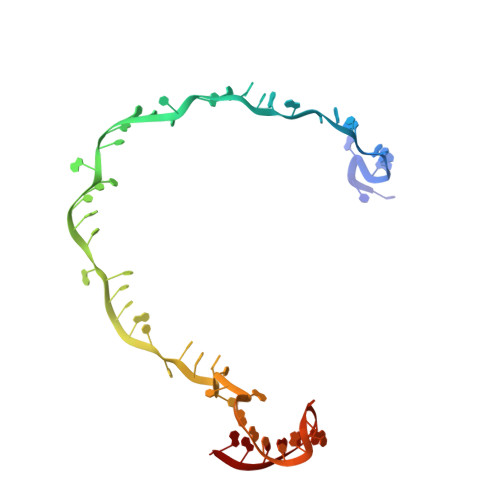
Structural basis for inhibition of the type I-F CRISPR-Cas surveillance complex by AcrIF4, AcrIF7 and AcrIF14.
Gabel, C., Li, Z., Zhang, H., Chang, L.(2021) Nucleic Acids Res 49: 584-594
- PubMed: 33332569
- DOI: https://doi.org/10.1093/nar/gkaa1199
- Primary Citation of Related Structures:
7JZW, 7JZX, 7JZZ - PubMed Abstract:
CRISPR-Cas systems are adaptive immune systems in bacteria and archaea to defend against mobile genetic elements (MGEs) and have been repurposed as genome editing tools. Anti-CRISPR (Acr) proteins are produced by MGEs to counteract CRISPR-Cas systems and can be used to regulate genome editing by CRISPR techniques. Here, we report the cryo-EM structures of three type I-F Acr proteins, AcrIF4, AcrIF7 and AcrIF14, bound to the type I-F CRISPR-Cas surveillance complex (the Csy complex) from Pseudomonas aeruginosa. AcrIF4 binds to an unprecedented site on the C-terminal helical bundle of Cas8f subunit, precluding conformational changes required for activation of the Csy complex. AcrIF7 mimics the PAM duplex of target DNA and is bound to the N-terminal DNA vise of Cas8f. Two copies of AcrIF14 bind to the thumb domains of Cas7.4f and Cas7.6f, preventing hybridization between target DNA and the crRNA. Our results reveal structural detail of three AcrIF proteins, each binding to a different site on the Csy complex for inhibiting degradation of MGEs.
Organizational Affiliation:
Department of Biological Sciences, Purdue University, 915 W. State Street, West Lafayette, IN 47907, USA.