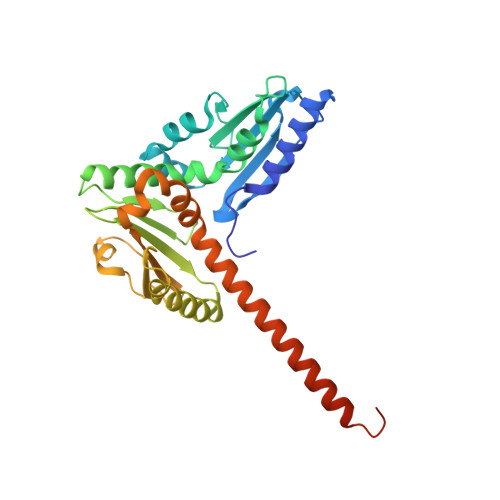
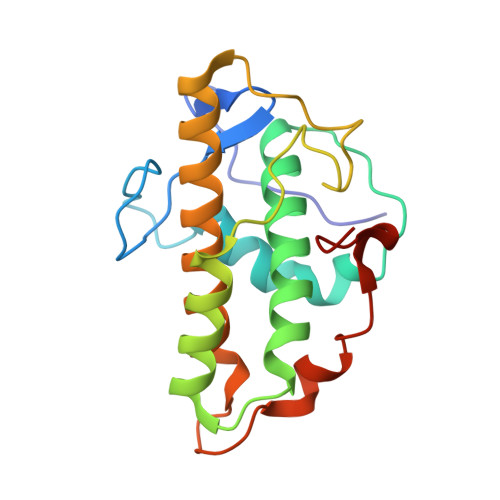
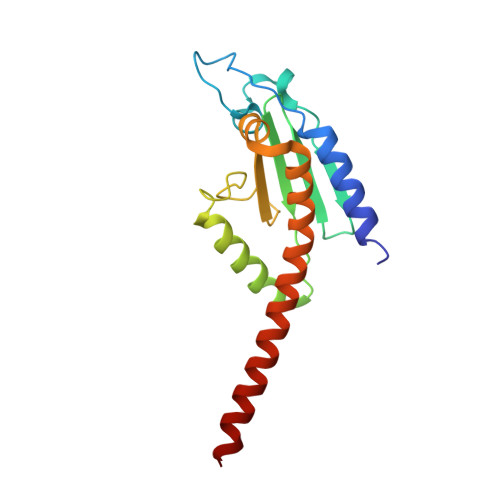
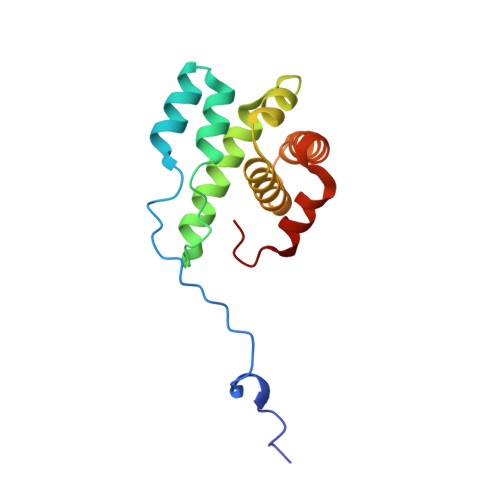
Molecular mechanism of Arp2/3 complex inhibition by Arpin.
Fregoso, F.E., van Eeuwen, T., Simanov, G., Rebowski, G., Boczkowska, M., Zimmet, A., Gautreau, A.M., Dominguez, R.(2022) Nat Commun 13: 628-628
- PubMed: 35110533
- DOI: https://doi.org/10.1038/s41467-022-28112-2
- Primary Citation of Related Structures:
7JPN - PubMed Abstract:
Positive feedback loops involving signaling and actin assembly factors mediate the formation and remodeling of branched actin networks in processes ranging from cell and organelle motility to mechanosensation. The Arp2/3 complex inhibitor Arpin controls the directional persistence of cell migration by interrupting a feedback loop involving Rac-WAVE-Arp2/3 complex, but Arpin's mechanism of inhibition is unknown. Here, we describe the cryo-EM structure of Arpin bound to Arp2/3 complex at 3.24-Å resolution. Unexpectedly, Arpin binds Arp2/3 complex similarly to WASP-family nucleation-promoting factors (NPFs) that activate the complex. However, whereas NPFs bind to two sites on Arp2/3 complex, on Arp2-ArpC1 and Arp3, Arpin only binds to the site on Arp3. Like NPFs, Arpin has a C-helix that binds at the barbed end of Arp3. Mutagenesis studies in vitro and in cells reveal how sequence differences within the C-helix define the molecular basis for inhibition by Arpin vs. activation by NPFs.
Organizational Affiliation:
Department of Physiology and Biochemistry and Molecular Biophysics Graduate Group, Perelman School of Medicine, University of Pennsylvania, Philadelphia, PA, 19104, USA.