The nitrosoamphetamine metabolite is accommodated in the active site of human hemoglobin: Spectroscopy and crystal structure.
Powell, S.M., Thomas, L.M., Richter-Addo, G.B.(2020) J Inorg Biochem 213: 111262-111262
- PubMed: 33049600
- DOI: https://doi.org/10.1016/j.jinorgbio.2020.111262
- Primary Citation of Related Structures:
7JJQ - PubMed Abstract:
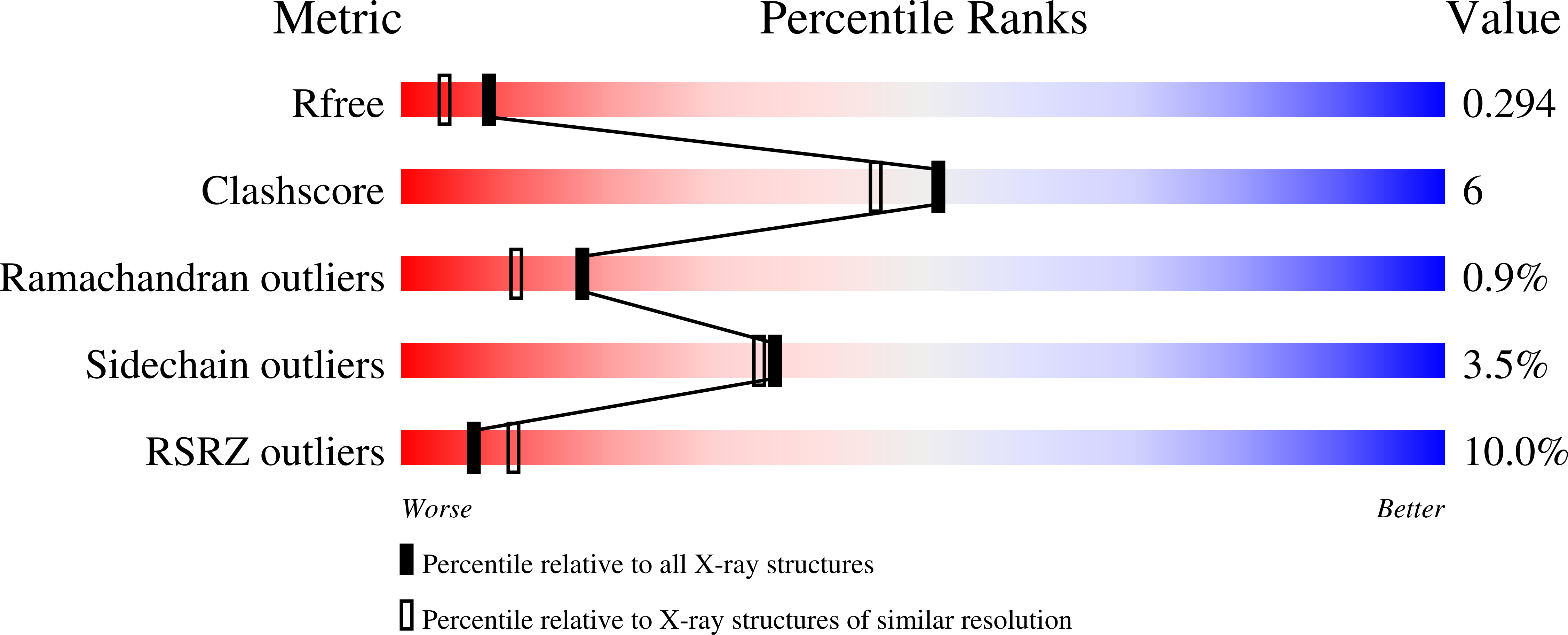
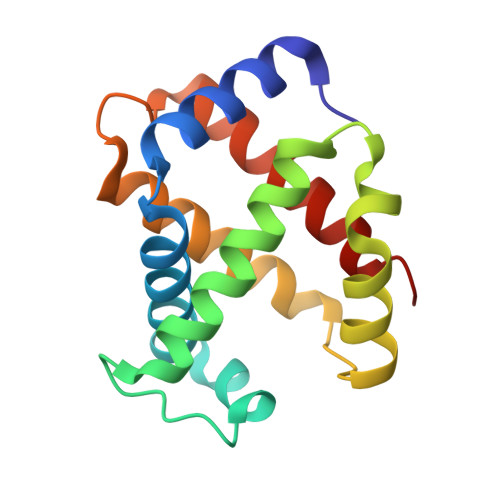
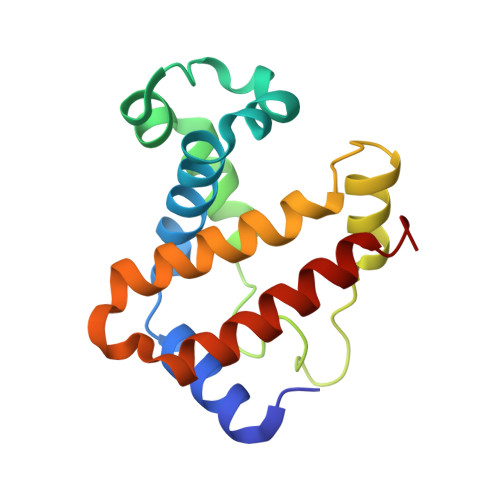
Amphetamine-based (Amph) drugs are metabolized in humans to their hydroxylamine (AmphNHOH) and nitroso (AmphNO) derivatives. The latter metabolites are known to bind to the Fe centers of cytochrome P450 and other heme enzymes to inhibit their activities. Although these AmphNHOH/AmphNO metabolites are present in vivo, their interactions with the blood protein hemoglobin (Hb) and the muscle protein (Mb) have been largely discounted due to a perception that the relatively small heme active sites of Hb and Mb will not be able to accommodate the large AmphNO group. We report the 2.15 Å resolution X-ray crystal structure of the AmphNO adduct of adult human hemoglobin as the Hb [α-Fe III (H 2 O)][β-Fe II (AmphNO)] derivative. We show that the binding of AmphNO to the β subunit is enabled by an E helix movement and stabilization of ligand binding by H-bonding with the distal His63 residue. We also observe an AmphNHOH group in the Xe2 pocket in close proximity to the α heme site in this derivative. Additionally, UV-vis spectroscopy was used to characterize this and related wt and mutant Mb adducts. Importantly, our X-ray crystal structure of this Hb-nitrosoamphetamine complex represents the first crystal structure of a wild-type heme protein adduct of any amphetamine metabolite. Our results provide a framework for further studies of AmphNHOH/AmphNO interactions with Hb and Mb as viable processes that potentially contribute to the overall biological inorganic chemistry of amphetamine drugs.
Organizational Affiliation:
Price Family Foundation Institute of Structural Biology, and Department of Chemistry and Biochemistry, University of Oklahoma, 101 Stephenson Parkway, Norman, OK 73019, United States of America.