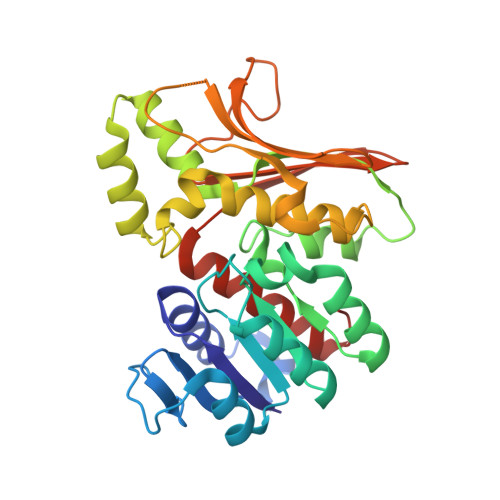
Conformational changes in the catalytic region are responsible for heat-induced activation of hyperthermophilic homoserine dehydrogenase.
Kubota, T., Kurihara, E., Watanabe, K., Ogata, K., Kaneko, R., Goto, M., Ohshima, T., Yoshimune, K.(2022) Commun Biol 5: 704-704
- PubMed: 35835834
- DOI: https://doi.org/10.1038/s42003-022-03656-7
- Primary Citation of Related Structures:
7F4B, 7F4C - PubMed Abstract:
When overexpressed as an immature enzyme in the mesophilic bacterium Escherichia coli, recombinant homoserine dehydrogenase from the hyperthermophilic archaeon Sulfurisphaera tokodaii (StHSD) was markedly activated by heat treatment. Both the apo- and holo-forms of the immature enzyme were successively crystallized, and the two structures were determined. Comparison among the structures of the immature enzyme and previously reported structures of mature enzymes revealed that a conformational change in a flexible part (residues 160-190) of the enzyme, which encloses substrates within the substrate-binding pocket, is smaller in the immature enzyme. The immature enzyme, but not the mature enzyme, formed a complex that included NADP + , despite its absence during crystallization. This indicates that the opening to the substrate-binding pocket in the immature enzyme is not sufficient for substrate-binding, efficient catalytic turnover or release of NADP + . Thus, specific conformational changes within the catalytic region appear to be responsible for heat-induced activation.
Organizational Affiliation:
Department of Applied Molecular Chemistry, Graduate School of Industrial Technology, Nihon University, 1-2-1, Izumichou, Narashino, Chiba, 275-8575, Japan.