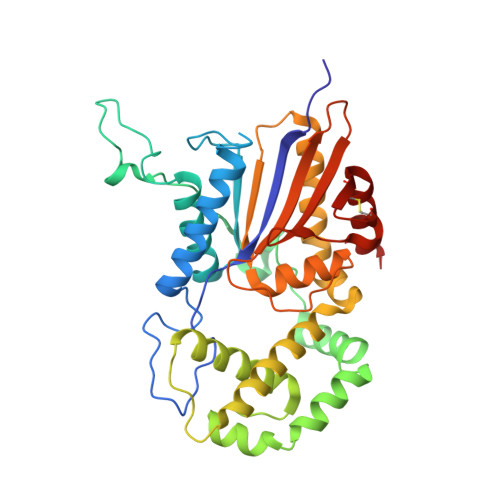
Structural insights into a new substrate binding mode of a histidine acid phosphatase from Legionella pneumophila.
Guo, Y., Zhou, D., Zhang, H., Zhang, N.N., Qi, X., Chen, X., Chen, Q., Li, J., Ge, H., Teng, Y.B.(2021) Biochem Biophys Res Commun 540: 90-94
- PubMed: 33450485
- DOI: https://doi.org/10.1016/j.bbrc.2020.12.070
- Primary Citation of Related Structures:
7D2F, 7DOQ - PubMed Abstract:
MapA is a histidine acid phosphatase (HAP) from Legionella pneumophila that catalyzes the hydroxylation of a phosphoryl group from phosphomonoesters by an active-site histidine. Several structures of HAPs, including MapA, in complex with the inhibitor tartrate have been solved and the substrate binding tunnel identified; however, the substrate recognition mechanism remains unknown. To gain insight into the mechanism of substrate recognition, the crystal structures of apo-MapA and the MapA D281A mutant in complex with 5'-AMP were solved at 2.2 and 2.6 Å resolution, respectively. The structure of the MapA D281A /5'-AMP complex reveals that the 5'-AMP fits fully into the substrate binding tunnel, with the 2'-hydroxyl group of the ribose moiety stabilized by Glu201 and the adenine moiety sandwiched between His205 and Phe237. This is the second structure of a HAP/AMP complex solved with 5'-AMP binding in a unique manner in the active site. The structure presents a new substrate recognition mechanism of HAPs.
Organizational Affiliation:
School of Life Sciences, Anhui Medical University, Hefei, Anhui, 230032, People's Republic of China. Electronic address: guoyu_joy@163.com.