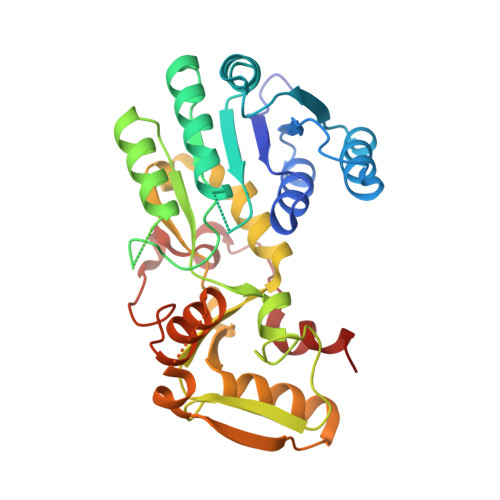
Structure-based engineering of substrate specificity for pinoresinol-lariciresinol reductases.
Xiao, Y., Shao, K., Zhou, J., Wang, L., Ma, X., Wu, D., Yang, Y., Chen, J., Feng, J., Qiu, S., Lv, Z., Zhang, L., Zhang, P., Chen, W.(2021) Nat Commun 12: 2828-2828
- PubMed: 33990581
- DOI: https://doi.org/10.1038/s41467-021-23095-y
- Primary Citation of Related Structures:
7CS2, 7CS3, 7CS4, 7CS5, 7CS6, 7CS7, 7CS8, 7CS9, 7CSA, 7CSB, 7CSC, 7CSD, 7CSE, 7CSF, 7CSG, 7CSH - PubMed Abstract:
Pinoresinol-lariciresinol reductases (PLRs) are enzymes involved in the lignan biosynthesis after the initial dimerization of two monolignols, and this represents the entry point for the synthesis of 8-8' lignans and contributes greatly to their structural diversity. Of particular interest has been the determination of how differing substrate specificities are achieved with these enzymes. Here, we present crystal structures of IiPLR1 from Isatis indigotica and pinoresinol reductases (PrRs) AtPrR1 and AtPrR2 from Arabidopsis thaliana, in the apo, substrate-bound and product-bound states. Each structure contains a head-to-tail homodimer, and the catalytic pocket comprises structural elements from both monomers. β4 loop covers the top of the pocket, and residue 98 from the loop governs catalytic specificity. The substrate specificities of IiPLR1 and AtPrR2 can be switched via structure-guided mutagenesis. Our study provides insight into the molecular mechanism underlying the substrate specificity of PLRs/PrRs and suggests an efficient strategy for the large-scale commercial production of the pharmaceutically valuable compound lariciresinol.
Organizational Affiliation:
Research and Development Center of Chinese Medicine Resources and Biotechnology, Institute of Chinese Materia Medica, Shanghai University of Traditional Chinese Medicine, Shanghai, China.