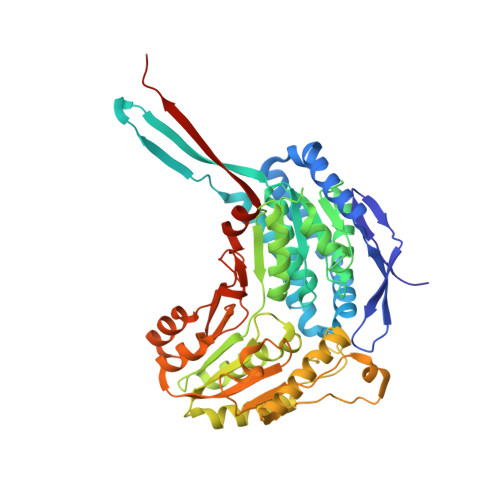
The tetrameric assembly of 2-aminomuconic 6-semialdehyde dehydrogenase is a functional requirement of cofactor NAD + binding.
Shi, Q., Chen, Y., Li, X., Dong, H., Chen, C., Zhong, Z., Yang, C., Liu, G., Su, D.(2021) Environ Microbiol
- PubMed: 34806815
- DOI: https://doi.org/10.1111/1462-2920.15840
- Primary Citation of Related Structures:
7BZV - PubMed Abstract:
The bacterium Pseudomonas sp. AP-3 is able to use the environmental pollutant 2-aminophenol as its sole source of carbon, nitrogen, and energy. Eight genes (amnA, B, C, D, E, F, G, and H) encoding 2-aminophenol metabolizing enzymes are clustered into a single operon. 2-Aminomuconic 6-semialdehyde dehydrogenase (AmnC), a member of the aldehyde dehydrogenase (ALDH) superfamily, is responsible for oxidizing 2-aminomuconic 6-semialdehyde to 2-aminomuconate. In contrast to many other members of the ALDH superfamily, the structural basis of the catalytic activity of AmnC remains elusive. Here, we present the crystal structure of AmnC, which displays a homotetrameric quaternary assembly that is directly involved in its enzymatic activity. The tetrameric state of AmnC in solution was also presented using small-angle X-ray scattering. The tetramerization of AmnC is mediated by the assembly of a protruding hydrophobic beta-strand motif and residues V121 and S123 located in the NAD + -binding domain of each subunit. Dimeric mutants of AmnC dramatically lose NAD + binding affinity and failed to oxidize the substrate analogue 2-hydroxymuconate-6-semialdehyde to α-hydroxymuconic acid, indicating that tetrameric assembly of AmnC is functional requirement.
Organizational Affiliation:
State Key Laboratory of Biotherapy, West China Hospital, Sichuan University, and Collaborative Innovation Center for Biotherapy, Chengdu, Sichuan, 610041, China.