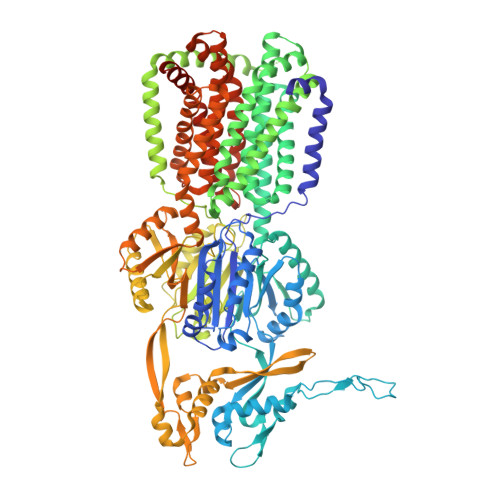
Structural and functional analysis of the promiscuous AcrB and AdeB efflux pumps suggests different drug binding mechanisms.
Ornik-Cha, A., Wilhelm, J., Kobylka, J., Sjuts, H., Vargiu, A.V., Malloci, G., Reitz, J., Seybert, A., Frangakis, A.S., Pos, K.M.(2021) Nat Commun 12: 6919-6919
- PubMed: 34824229
- DOI: https://doi.org/10.1038/s41467-021-27146-2
- Primary Citation of Related Structures:
7B8P, 7B8Q, 7B8R, 7B8S, 7B8T - PubMed Abstract:
Upon antibiotic stress Gram-negative pathogens deploy resistance-nodulation-cell division-type tripartite efflux pumps. These include a H + /drug antiporter module that recognizes structurally diverse substances, including antibiotics. Here, we show the 3.5 Å structure of subunit AdeB from the Acinetobacter baumannii AdeABC efflux pump solved by single-particle cryo-electron microscopy. The AdeB trimer adopts mainly a resting state with all protomers in a conformation devoid of transport channels or antibiotic binding sites. However, 10% of the protomers adopt a state where three transport channels lead to the closed substrate (deep) binding pocket. A comparison between drug binding of AdeB and Escherichia coli AcrB is made via activity analysis of 20 AdeB variants, selected on basis of side chain interactions with antibiotics observed in the AcrB periplasmic domain X-ray co-structures with fusidic acid (2.3 Å), doxycycline (2.1 Å) and levofloxacin (2.7 Å). AdeABC, compared to AcrAB-TolC, confers higher resistance to E. coli towards polyaromatic compounds and lower resistance towards antibiotic compounds.
Organizational Affiliation:
Institute of Biochemistry, Goethe-University Frankfurt, Max-von-Laue-Straße 9, 60438, Frankfurt am Main, Germany.