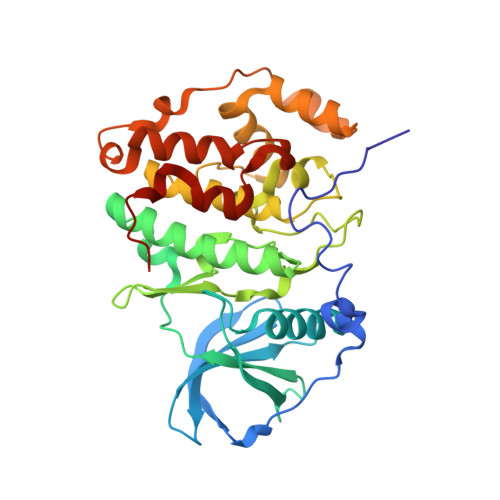
Structural basis for the design of bisubstrate inhibitors of protein kinase CK2 provided by complex structures with the substrate-competitive inhibitor heparin.
Schnitzler, A., Niefind, K.(2021) Eur J Med Chem 214: 113223-113223
- PubMed: 33571828
- DOI: https://doi.org/10.1016/j.ejmech.2021.113223
- Primary Citation of Related Structures:
7B8H, 7B8I - PubMed Abstract:
The Ser/Thr kinase CK2, a member of the superfamily of eukaryotic protein kinases, has an acidophilic substrate profile with the substrate recognition sequence S/T-D/E-X-D/E, and it is inhibited by polyanionic substances like heparin. The latter, a highly sulphated glucosamino glycan composed mainly of repeating 2-O-sulpho-α-l-idopyranuronic acid/N,O6-disulpho-α-d-glucosamine disaccharide units, is the longest known substrate-competitive CK2 inhibitor. The structural basis of CK2's preference for anionic substrates and substrate-competitive inhibitors is only vaguely known which limits the value of the substrate-binding region for the structure-based development of CK2 bisubstrate inhibitors. Here, a tetragonal and a monoclinic co-crystal structure of CK2α, the catalytic subunit of CK2, with a decameric heparin fragment are described. In the tetragonal structure, the heparin molecule binds to the polybasic stretch at the beginning of CK2α's helix αC, whereas in the monoclinic structure it occupies the central substrate-recognition region around the P+1 loop. Together, the structures rationalize the inhibitory efficacy of heparin fragments as a function of chain length. The monoclinic CK2α/heparin structure, in which the heparin fragment is particularly well defined, is the first CK2 structure with an anionic inhibitor of considerable size at the central part of the substrate-recognition site. The bound heparin fragment is so close to the binding site of ATP-competitive inhibitors that it can guide the design of linkers and pave the way to efficient CK2 bisubstrate inhibitors in the future.
Organizational Affiliation:
Universität zu Köln, Department für Chemie, Institut für Biochemie, Zülpicher Straße 47, D-50674 Köln, Germany.