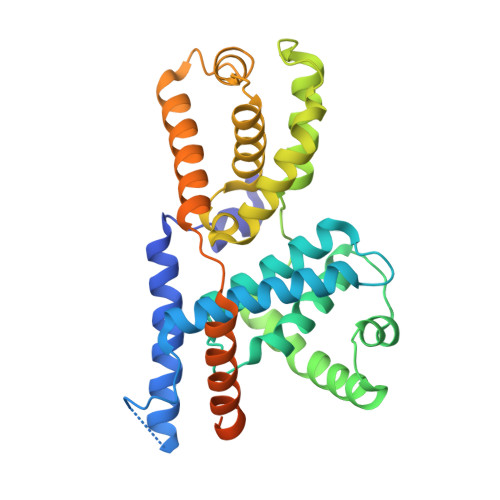
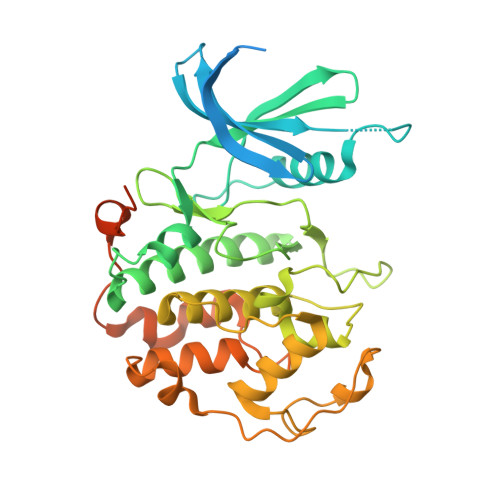
2.5 angstrom -resolution structure of human CDK-activating kinase bound to the clinical inhibitor ICEC0942.
Greber, B.J., Remis, J., Ali, S., Nogales, E.(2021) Biophys J 120: 677-686
- PubMed: 33476598
- DOI: https://doi.org/10.1016/j.bpj.2020.12.030
- Primary Citation of Related Structures:
7B5O, 7B5Q - PubMed Abstract:
The human CDK-activating kinase (CAK), composed of CDK7, cyclin H, and MAT1, is involved in the control of transcription initiation and the cell cycle. Because of these activities, it has been identified as a promising target for cancer chemotherapy. A number of CDK7 inhibitors have entered clinical trials, among them ICEC0942 (also known as CT7001). Structural information can aid in improving the affinity and specificity of such drugs or drug candidates, reducing side effects in patients. Here, we have determined the structure of the human CAK in complex with ICEC0942 at 2.5 Å-resolution using cryogenic electron microscopy. Our structure reveals conformational differences of ICEC0942 compared with previous X-ray crystal structures of the CDK2-bound complex, and highlights the critical ability of cryogenic electron microscopy to resolve structures of drug-bound protein complexes without the need to crystalize the protein target.
Organizational Affiliation:
Division of Structural Biology, The Institute of Cancer Research, London, United Kingdom; California Institute for Quantitative Biosciences (QB3), University of California, Berkeley, Berkeley, California; Molecular Biophysics and Integrative Bio-Imaging Division, Lawrence Berkeley National Laboratory, Berkeley, California. Electronic address: basil.greber@icr.ac.uk.