X-RAY CRYSTAL STRUCTURE OF THE CsPYL1-Lig2-HAB1 TERNARY COMPLEX
Albert, A., Infantes, L., Benavente, J.L.To be published.
Experimental Data Snapshot
Starting Model: experimental
View more details
Entity ID: 1 | |||||
|---|---|---|---|---|---|
| Molecule | Chains | Sequence Length | Organism | Details | Image |
| CsPYL1 | 209 | Citrus sinensis | Mutation(s): 0 Gene Names: CISIN_1g046151mg | 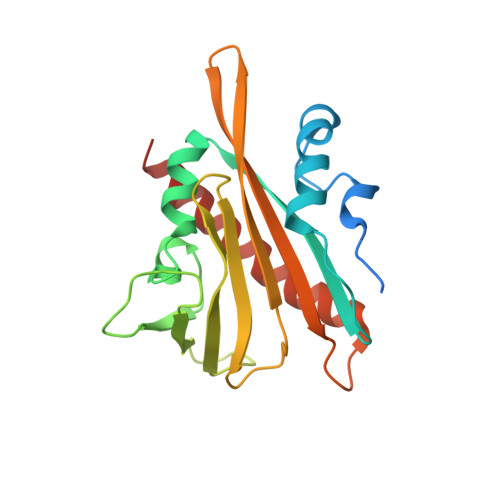 | |
UniProt | |||||
Find proteins for A0A067E666 (Citrus sinensis) Explore A0A067E666 Go to UniProtKB: A0A067E666 | |||||
Entity Groups | |||||
| Sequence Clusters | 30% Identity50% Identity70% Identity90% Identity95% Identity100% Identity | ||||
| UniProt Group | A0A067E666 | ||||
Sequence AnnotationsExpand | |||||
| |||||
Entity ID: 2 | |||||
|---|---|---|---|---|---|
| Molecule | Chains | Sequence Length | Organism | Details | Image |
| Protein phosphatase 2C 16 | 333 | Arabidopsis thaliana | Mutation(s): 0 Gene Names: HAB1, P2C-HA, At1g72770, F28P22.4 EC: 3.1.3.16 | 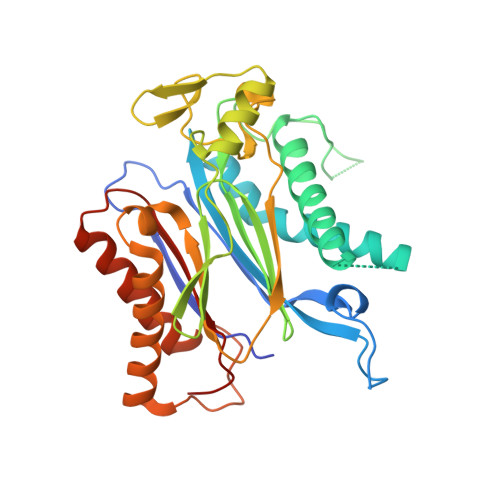 | |
UniProt | |||||
Find proteins for Q9CAJ0 (Arabidopsis thaliana) Explore Q9CAJ0 Go to UniProtKB: Q9CAJ0 | |||||
Entity Groups | |||||
| Sequence Clusters | 30% Identity50% Identity70% Identity90% Identity95% Identity100% Identity | ||||
| UniProt Group | Q9CAJ0 | ||||
Sequence AnnotationsExpand | |||||
| |||||
| Ligands 4 Unique | |||||
|---|---|---|---|---|---|
| ID | Chains | Name / Formula / InChI Key | 2D Diagram | 3D Interactions | |
| S5N (Subject of Investigation/LOI) Query on S5N | C [auth A] | N-((1,4-dimethyl-2-oxo-1,2-dihydroquinolin-6-yl)methyl)benzenesulfonamide C18 H18 N2 O3 S JYPYWRMUXLPCGO-UHFFFAOYSA-N |  | ||
| GOL Query on GOL | D [auth A], E [auth A], N [auth B] | GLYCEROL C3 H8 O3 PEDCQBHIVMGVHV-UHFFFAOYSA-N |  | ||
| MN Query on MN | F [auth B], G [auth B], H [auth B], I [auth B] | MANGANESE (II) ION Mn WAEMQWOKJMHJLA-UHFFFAOYSA-N |  | ||
| CL Query on CL | J [auth B], K [auth B], L [auth B], M [auth B] | CHLORIDE ION Cl VEXZGXHMUGYJMC-UHFFFAOYSA-M |  | ||
| Length ( Å ) | Angle ( ˚ ) |
|---|---|
| a = 43.118 | α = 90 |
| b = 62.45 | β = 90 |
| c = 187.578 | γ = 90 |
| Software Name | Purpose |
|---|---|
| PHENIX | refinement |
| XDS | data reduction |
| XDS | data scaling |
| PHASER | phasing |
| Funding Organization | Location | Grant Number |
|---|---|---|
| Ministry of Economy and Competitiveness (MINECO) | Spain | BIO2017-89523-R |