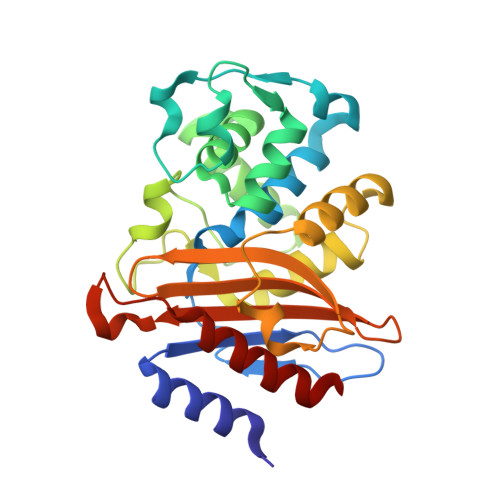
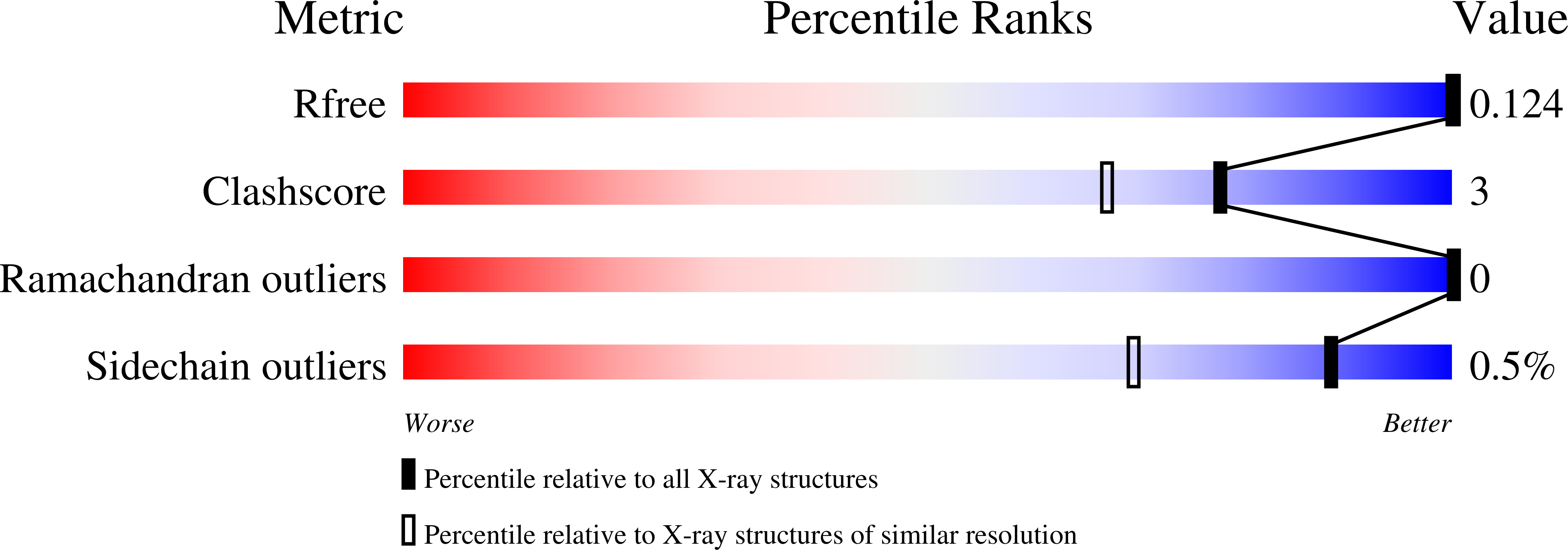Conserved proline residues prevent dimerization and aggregation in the beta-lactamase BlaC.
Chikunova, A., Manley, M.P., Heijjer, C.N., Drenth, C.S., Cramer-Blok, A.J., Ahmad, M.U.D., Perrakis, A., Ubbink, M.(2024) Protein Sci 33: e4972-e4972
- PubMed: 38533527
- DOI: https://doi.org/10.1002/pro.4972
- Primary Citation of Related Structures:
7A6Z, 8R88 - PubMed Abstract:
Evolution leads to conservation of amino acid residues in protein families. Conserved proline residues are usually considered to ensure the correct folding and to stabilize the three-dimensional structure. Surprisingly, proline residues that are highly conserved in class A β-lactamases were found to tolerate various substitutions without large losses in enzyme activity. We investigated the roles of three conserved prolines at positions 107, 226, and 258 in the β-lactamase BlaC from Mycobacterium tuberculosis and found that mutations can lead to dimerization of the enzyme and an overall less stable protein that is prone to aggregate over time. For the variant Pro107Thr, the crystal structure shows dimer formation resembling domain swapping. It is concluded that the proline substitutions loosen the structure, enhancing multimerization. Even though the enzyme does not lose its properties without the conserved proline residues, the prolines ensure the long-term structural integrity of the enzyme.
Organizational Affiliation:
Leiden Institute of Chemistry, Leiden University, Leiden, The Netherlands.