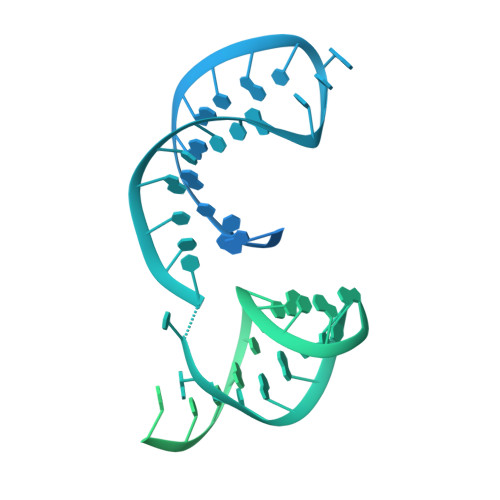
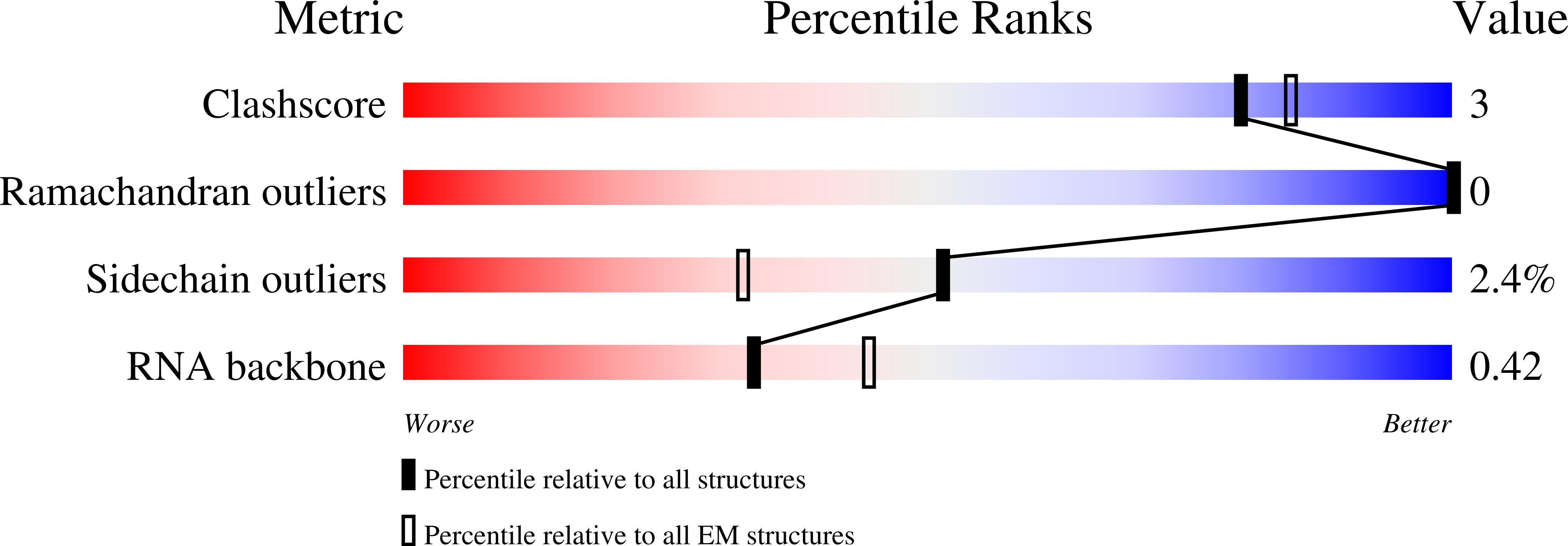
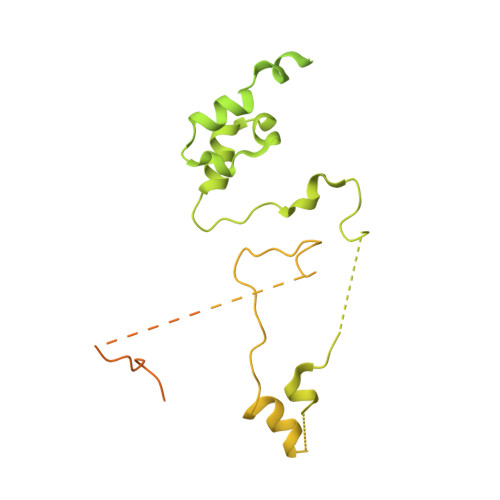
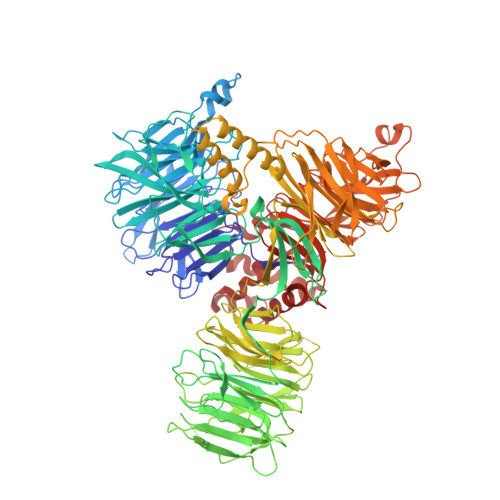
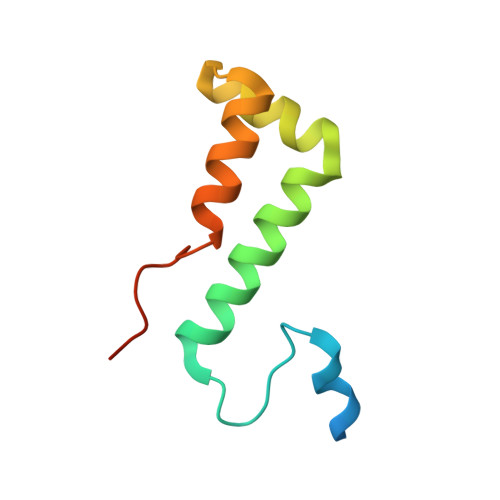
Structural basis of branch site recognition by the human spliceosome.
Tholen, J., Razew, M., Weis, F., Galej, W.P.(2022) Science 375: 50-57
- PubMed: 34822310
- DOI: https://doi.org/10.1126/science.abm4245
- Primary Citation of Related Structures:
7Q3L, 7Q4O, 7Q4P - PubMed Abstract:
Recognition of the intron branch site (BS) by the U2 small nuclear ribonucleoprotein (snRNP) is a critical event during spliceosome assembly. In mammals, BS sequences are poorly conserved, and unambiguous intron recognition cannot be achieved solely through a base-pairing mechanism. We isolated human 17 S U2 snRNP and reconstituted in vitro its adenosine 5´-triphosphate (ATP)–dependent remodeling and binding to the pre–messenger RNA substrate. We determined a series of high-resolution (2.0 to 2.2 angstrom) structures providing snapshots of the BS selection process. The substrate-bound U2 snRNP shows that SF3B6 stabilizes the BS:U2 snRNA duplex, which could aid binding of introns with poor sequence complementarity. ATP-dependent remodeling uncoupled from substrate binding captures U2 snRNA in a conformation that competes with BS recognition, providing a selection mechanism based on branch helix stability.
Organizational Affiliation:
European Molecular Biology Laboratory, 71 Avenue des Martyrs, 38042 Grenoble, France.