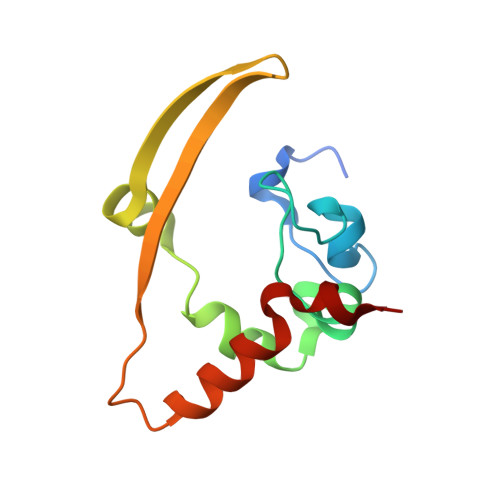
Structures of the SARS-CoV-2 nucleocapsid and their perspectives for drug design.
Peng, Y., Du, N., Lei, Y., Dorje, S., Qi, J., Luo, T., Gao, G.F., Song, H.(2020) EMBO J 39: e105938-e105938
- PubMed: 32914439
- DOI: https://doi.org/10.15252/embj.2020105938
- Primary Citation of Related Structures:
7CDZ, 7CE0 - PubMed Abstract:
COVID-19, caused by SARS-CoV-2, has resulted in severe and unprecedented economic and social disruptions in the world. Nucleocapsid (N) protein, which is the major structural component of the virion and is involved in viral replication, assembly and immune regulation, plays key roles in the viral life cycle. Here, we solved the crystal structures of the N- and C-terminal domains (N-NTD and N-CTD) of SARS-CoV-2 N protein, at 1.8 and 1.5 Å resolution, respectively. Both structures show conserved features from other CoV N proteins. The binding sites targeted by small molecules against HCoV-OC43 and MERS-CoV, which inhibit viral infection by blocking the RNA-binding activity or normal oligomerization of N protein, are relatively conserved in our structure, indicating N protein is a promising drug target. In addition, certain areas of N-NTD and N-CTD display distinct charge distribution patterns in SARS-CoV-2, which may alter the RNA-binding modes. The specific antigenic characteristics are critical for developing specific immune-based rapid diagnostic tests. Our structural information can aid in the discovery and development of antiviral inhibitors against SARS-CoV-2 in the future.
Organizational Affiliation:
Laboratory of Animal Infectious Diseases, College of Animal Sciences and Veterinary Medicine, Guangxi University, Nanning, China.