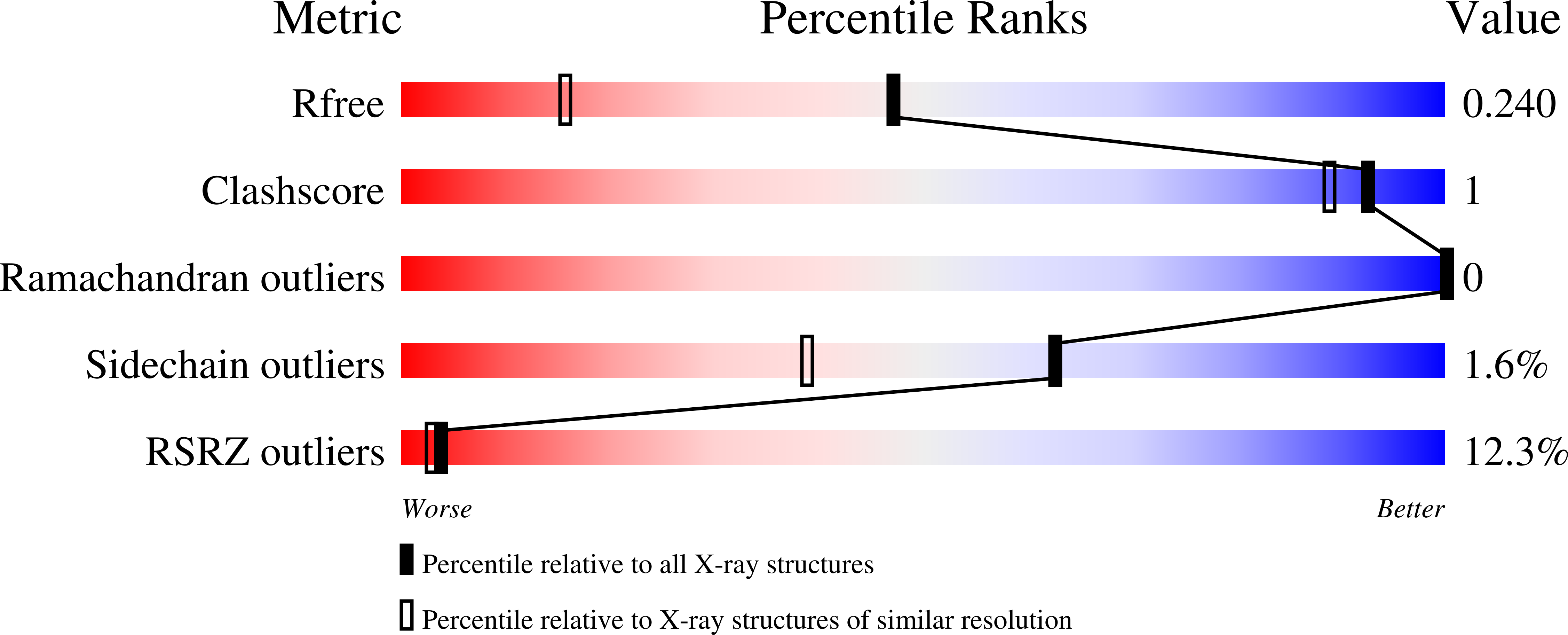
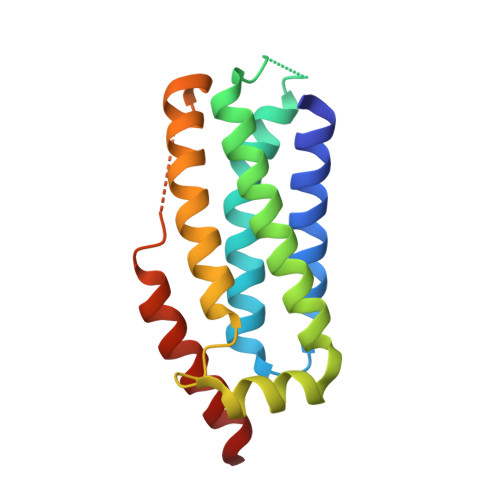
Locking mixed-lineage kinase domain-like protein in its auto-inhibited state prevents necroptosis.
Rubbelke, M., Fiegen, D., Bauer, M., Binder, F., Hamilton, J., King, J., Thamm, S., Nar, H., Zeeb, M.(2020) Proc Natl Acad Sci U S A 117: 33272-33281
- PubMed: 33318170
- DOI: https://doi.org/10.1073/pnas.2017406117
- Primary Citation of Related Structures:
6ZLE, 6ZPR, 6ZVO, 6ZZ1 - PubMed Abstract:
As an alternative pathway of controlled cell death, necroptosis can be triggered by tumor necrosis factor via the kinases RIPK1/RIPK3 and the effector protein mixed-lineage kinase domain-like protein (MLKL). Upon activation, MLKL oligomerizes and integrates into the plasma membrane via its executioner domain. Here, we present the X-ray and NMR costructures of the human MLKL executioner domain covalently bound via Cys86 to a xanthine class inhibitor. The structures reveal that the compound stabilizes the interaction between the auto-inhibitory brace helix α6 and the four-helix bundle by stacking to Phe148. An NMR-based functional assay observing the conformation of this helix showed that the F148A mutant is unresponsive to the compound, providing further evidence for the importance of this interaction. Real-time and diffusion NMR studies demonstrate that xanthine derivatives inhibit MLKL oligomerization. Finally, we show that the other well-known MLKL inhibitor Necrosulfonamide, which also covalently modifies Cys86, must employ a different mode of action.
Organizational Affiliation:
Medicinal Chemistry, Boehringer Ingelheim Pharma GmbH & Co. KG, 88397 Biberach an der Riss, Germany.