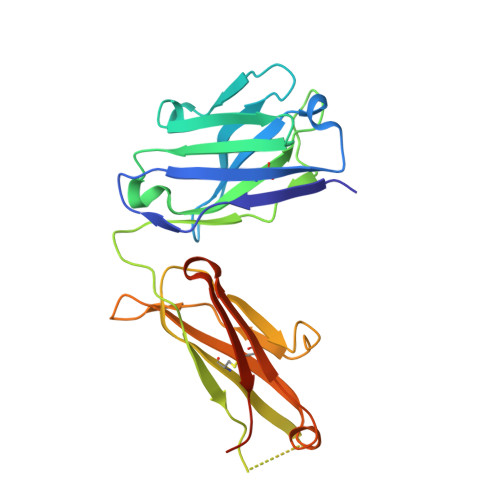
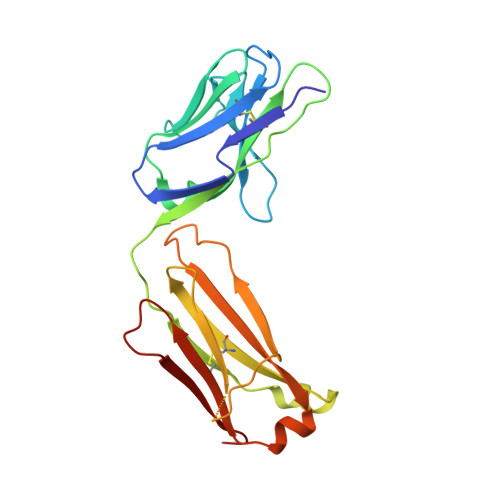
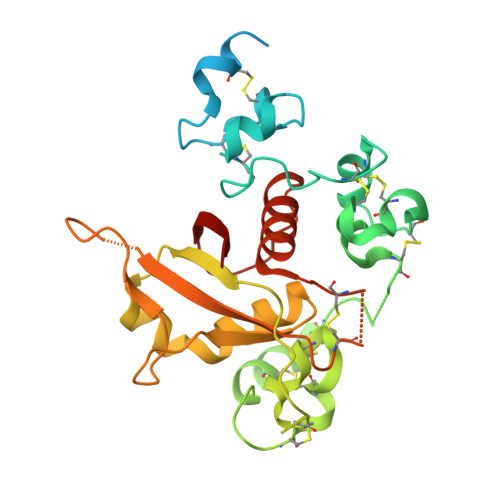
NOTCH3-targeted antibody drug conjugates regress tumors by inducing apoptosis in receptor cells and through transendocytosis into ligand cells.
Geles, K.G., Gao, Y., Giannakou, A., Sridharan, L., Yamin, T.T., Zhang, J., Karim, R., Bard, J., Piche-Nicholas, N., Charati, M., Maderna, A., Lucas, J., Golas, J., Guffroy, M., Pirie-Shepherd, S., Roy, M., Qian, J., Franks, T., Zhong, W., O'Donnell, C.J., Tchistiakova, L., Gerber, H.P., Sapra, P.(2021) Cell Rep Med 2: 100279-100279
- PubMed: 34095881
- DOI: https://doi.org/10.1016/j.xcrm.2021.100279
- Primary Citation of Related Structures:
6XSW - PubMed Abstract:
Aberrant NOTCH3 signaling and overexpression is oncogenic, associated with cancer stem cells and drug resistance, yet therapeutic targeting remains elusive. Here, we develop NOTCH3-targeted antibody drug conjugates (NOTCH3-ADCs) by bioconjugation of an auristatin microtubule inhibitor through a protease cleavable linker to two antibodies with differential abilities to inhibit signaling. The signaling inhibitory antibody rapidly induces ligand-independent receptor clustering and internalization through both caveolin and clathrin-mediated pathways. The non-inhibitory antibody also efficiently endocytoses via clathrin without inducing receptor clustering but with slower lysosomal co-localization kinetics. In addition, DLL4 ligand binding to the NOTCH3 receptor mediates transendocytosis of NOTCH3-ADCs into ligand-expressing cells. NOTCH3-ADCs internalize into receptor and ligand cells independent of signaling and induce cell death in both cell types representing an atypical mechanism of ADC cytotoxicity. Treatment of xenografts with NOTCH3-ADCs leads to sustained tumor regressions, outperforms standard-of-care chemotherapy, and allows targeting of tumors that overexpress NOTCH3 independent of signaling inhibition.
Organizational Affiliation:
Pfizer Worldwide Research and Development, Oncology Research and Development, Pearl River, NY, USA.