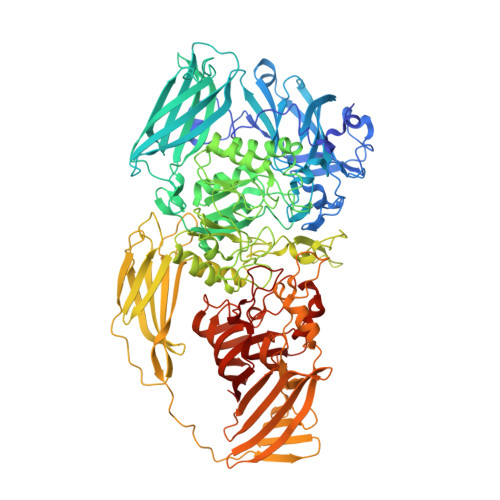
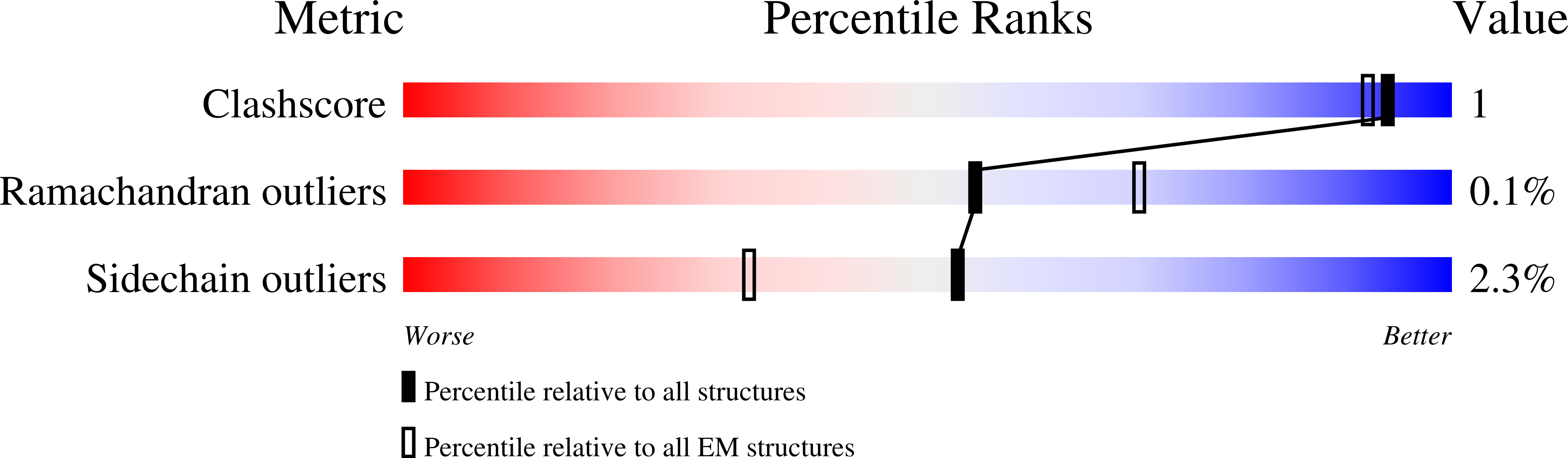Fragment-based drug discovery using cryo-EM.
Saur, M., Hartshorn, M.J., Dong, J., Reeks, J., Bunkoczi, G., Jhoti, H., Williams, P.A.(2020) Drug Discov Today 25: 485-490
- PubMed: 31877353
- DOI: https://doi.org/10.1016/j.drudis.2019.12.006
- Primary Citation of Related Structures:
6TSH, 6TSK, 6TTE, 6TTF, 6TTH, 6TTI, 6TTQ - PubMed Abstract:
Recent advances in electron cryo-microscopy (cryo-EM) structure determination have pushed the resolutions obtainable by the method into the range widely considered to be of utility for drug discovery. Here, we review the use of cryo-EM in fragment-based drug discovery (FBDD) based on in-house method development. We demonstrate not only that cryo-EM can reveal details of the molecular interactions between fragments and a protein, but also that the current reproducibility, quality, and throughput are compatible with FBDD. We exemplify this using the test system β-galactosidase (Bgal) and the oncology target pyruvate kinase 2 (PKM2).
Organizational Affiliation:
Astex Pharmaceuticals, 436 Cambridge Science Park, Cambridge, CB4 0QA, UK.