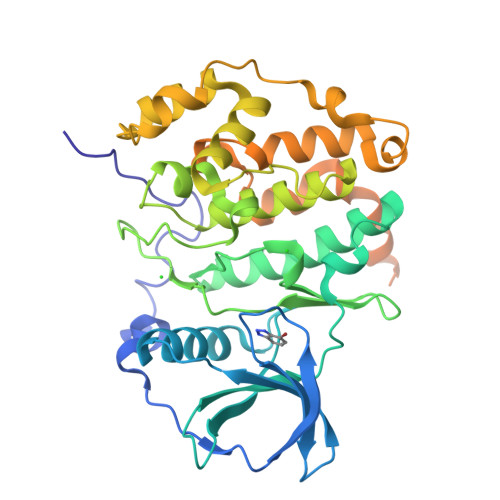
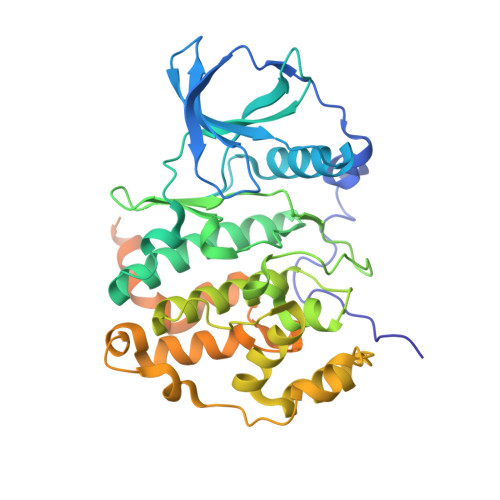
Halogen Atoms in the Protein-Ligand System. Structural and Thermodynamic Studies of the Binding of Bromobenzotriazoles by the Catalytic Subunit of Human Protein Kinase CK2.
Czapinska, H., Winiewska-Szajewska, M., Szymaniec-Rutkowska, A., Piasecka, A., Bochtler, M., Poznanski, J.(2021) J Phys Chem B 125: 2491-2503
- PubMed: 33689348
- DOI: https://doi.org/10.1021/acs.jpcb.0c10264
- Primary Citation of Related Structures:
6TLL, 6TLO, 6TLP, 6TLR, 6TLS, 6TLU, 6TLV, 6TLW - PubMed Abstract:
Binding of a family of brominated benzotriazoles to the catalytic subunit of human protein kinase CK2 (hCK2α) was used as a model system to assess the contribution of halogen bonding to protein-ligand interaction. CK2 is a constitutively active pleiotropic serine/threonine protein kinase that belongs to the CMGC group of eukaryotic protein kinases (EPKs). Due to the addiction of some cancer cells, CK2 is an attractive and well-characterized drug target. Halogenated benzotriazoles act as ATP-competitive inhibitors with unexpectedly good selectivity for CK2 over other EPKs. We have characterized the interaction of bromobenzotriazoles with hCK2α by X-ray crystallography, low-volume differential scanning fluorimetry, and isothermal titration calorimetry. Properties of free ligands in solution were additionally characterized by volumetric and RT-HPLC measurements. Thermodynamic data indicate that the affinity increases with bromo substitution, with greater contributions from 5- and 6-substituents than 4- and 7-substituents. Except for 4,7-disubstituted compounds, the bromobenzotriazoles adopt a canonical pose with the triazole close to lysine 68, which precludes halogen bonding. More highly substituted benzotriazoles adopt many additional noncanonical poses, presumably driven by a large hydrophobic contribution to binding. Some noncanonical ligand orientations allow the formation of halogen bonds with the hinge region. Consistent with a predominantly hydrophobic interaction, the isobaric heat capacity decreases upon ligand binding, the more so the higher the substitution.
Organizational Affiliation:
Institute of Biochemistry and Biophysics PAS, Pawińskiego 5a, 02-106 Warsaw, Poland.