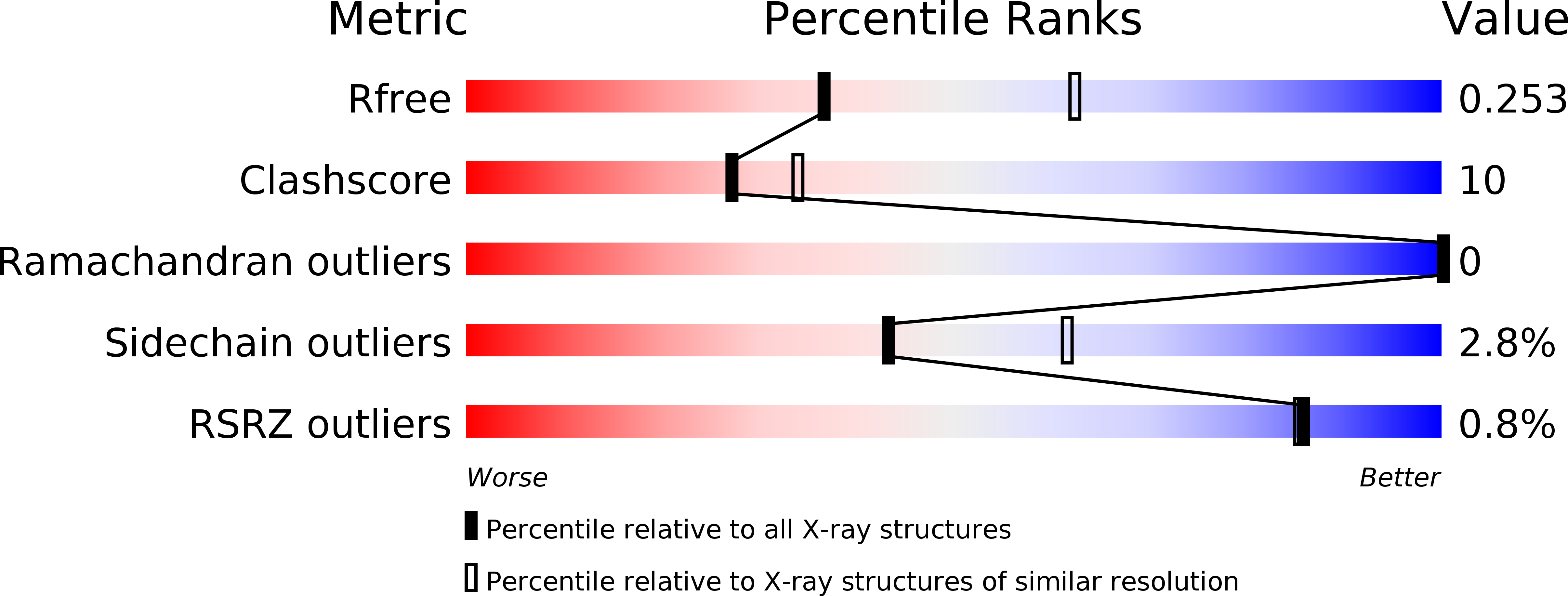
A FabG inhibitor targeting an allosteric binding site inhibits several orthologs from Gram-negative ESKAPE pathogens.
Vella, P., Rudraraju, R.S., Lundback, T., Axelsson, H., Almqvist, H., Vallin, M., Schneider, G., Schnell, R.(2021) Bioorg Med Chem 30: 115898-115898
- PubMed: 33388594
- DOI: https://doi.org/10.1016/j.bmc.2020.115898
- Primary Citation of Related Structures:
6T5X, 6T60, 6T62, 6T65, 6T6N, 6T6P, 6T77, 6T7M - PubMed Abstract:
The spread of antibiotic resistance within the ESKAPE group of human pathogenic bacteria poses severe challenges in the treatment of infections and maintenance of safe hospital environments. This motivates efforts to validate novel target proteins within these species that could be pursued as potential targets for antibiotic development. Genetic data suggest that the enzyme FabG, which is part of the bacterial fatty acid biosynthetic system FAS-II, is essential in several ESKAPE pathogens. FabG catalyzes the NADPH dependent reduction of 3-keto-acyl-ACP during fatty acid elongation, thus enabling lipid supply for production and maintenance of the cell envelope. Here we report on small-molecule screening on the FabG enzymes from A. baumannii and S. typhimurium to identify a set of µM inhibitors, with the most potent representative (1) demonstrating activity against six FabG-orthologues. A co-crystal structure with FabG from A. baumannii (PDB:6T65) confirms inhibitor binding at an allosteric site located in the subunit interface, as previously demonstrated for other sub-µM inhibitors of FabG from P. aeruginosa. We show that inhibitor binding distorts the oligomerization interface in the FabG tetramer and displaces crucial residues involved in the interaction with the co-substrate NADPH. These observations suggest a conserved allosteric site across the FabG family, which can be potentially targeted for interference with fatty acid biosynthesis in clinically relevant ESKAPE pathogens.
Organizational Affiliation:
Department of Medical Biochemistry and Biophysics, Karolinska Institutet, S-17 165 Stockholm, Sweden.