Design, Synthesis, Evaluation, and Structural Studies ofC2-Symmetric Small Molecule Inhibitors of Programmed Cell Death-1/Programmed Death-Ligand 1 Protein-Protein Interaction.
Basu, S., Yang, J., Xu, B., Magiera-Mularz, K., Skalniak, L., Musielak, B., Kholodovych, V., Holak, T.A., Hu, L.(2019) J Med Chem 62: 7250-7263
- PubMed: 31298541
- DOI: https://doi.org/10.1021/acs.jmedchem.9b00795
- Primary Citation of Related Structures:
6RPG - PubMed Abstract:
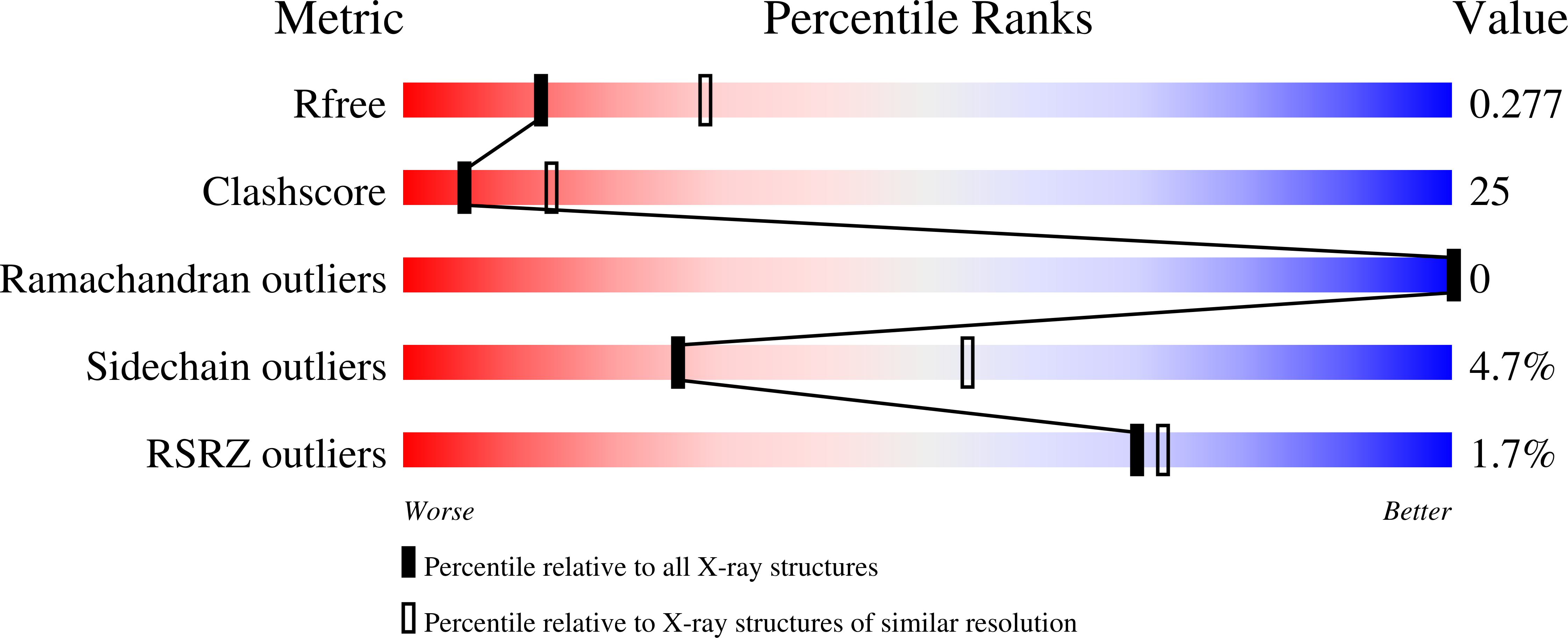
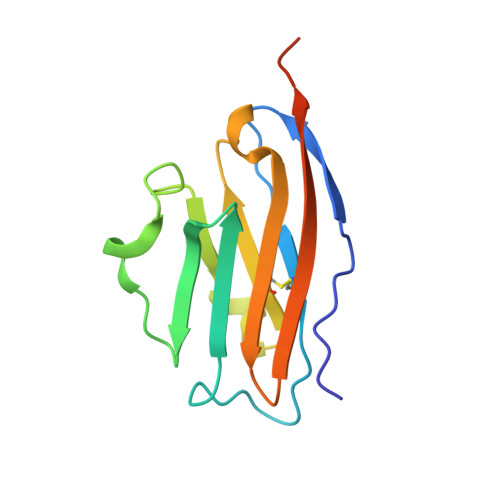
A series of C 2 -symmetric inhibitors was designed and evaluated for inhibitory activity against the programmed cell death-1/programmed death-ligand 1(PD-1/PD-L1) protein-protein interaction (PPI) in a homogenous time-resolved fluorescence (HTRF) assay and PD-1 signaling in cell-based coculture assays. C 2 -symmetric inhibitors 2a (LH1306) and 2b (LH1307) exhibited IC 50 values of 25 and 3.0 nM, respectively, in the HTRF assay. While 2a was ∼3.8-fold more potent than previously reported inhibitor 1a , 2b could not be differentiated from 1b due to their high potency and the limit of our HTRF assay conditions. In one cell-based coculture PD-1 signaling assay, 2a and 2b were 8.2- and 2.8-fold more potent in inhibiting PD-1 signaling than 1a and 1b , respectively. NMR and X-ray cocrystal structural studies provided more structural insights into the interaction between 2b and PD-L1; 2b binds to PD-L1 at the PD-1 binding site and induces the formation of a more symmetrically arranged PD-L1 homodimer than that previously reported for other inhibitors.
Organizational Affiliation:
Department of Organic Chemistry, Faculty of Chemistry , Jagiellonian University , Gronostajowa 2 , 30-387 Krakow , Poland.