The characterization of Thermotoga maritima Arginine Binding Protein variants demonstrates that minimal local strains have an important impact on protein stability.
Balasco, N., Smaldone, G., Vigorita, M., Del Vecchio, P., Graziano, G., Ruggiero, A., Vitagliano, L.(2019) Sci Rep 9: 6617-6617
- PubMed: 31036855
- DOI: https://doi.org/10.1038/s41598-019-43157-y
- Primary Citation of Related Structures:
6Q3U - PubMed Abstract:
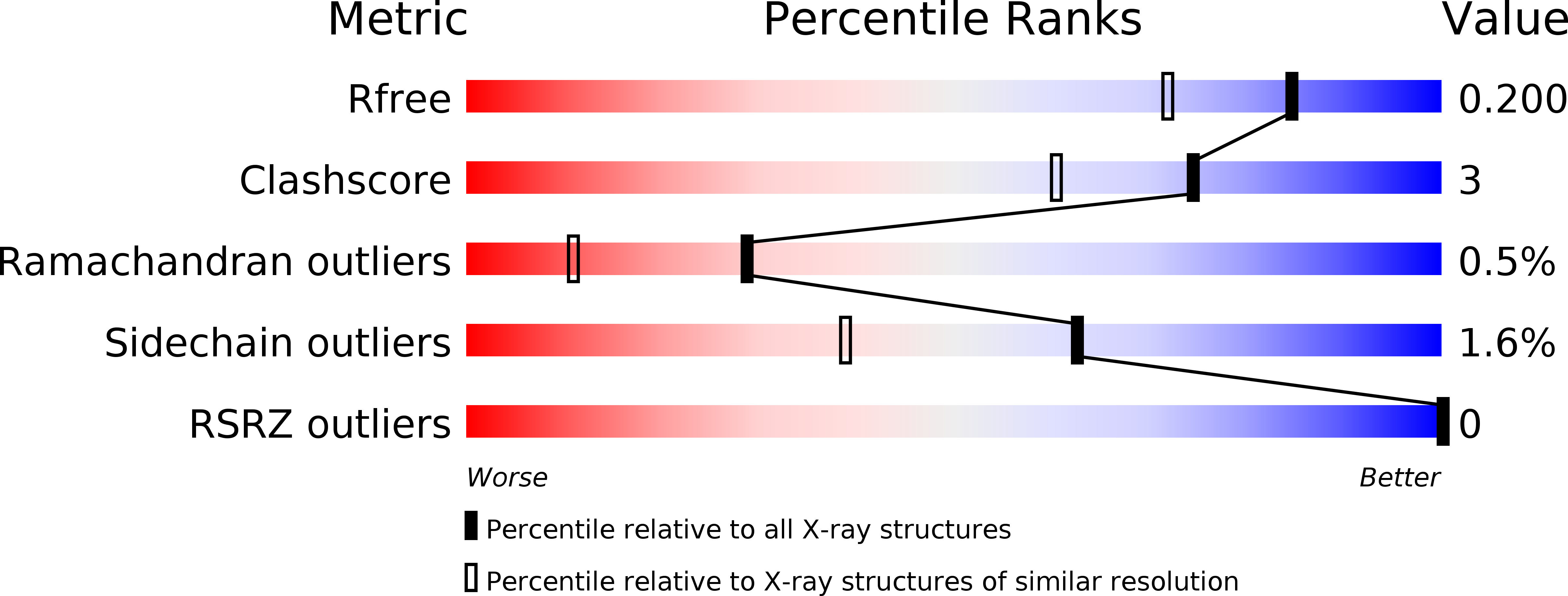
The Ramachandran plot is a versatile and valuable tool that provides fundamental information for protein structure determination, prediction, and validation. The structural/thermodynamic effects produced by forcing a residue to adopt a conformation predicted to be forbidden were here explored using Thermotoga maritima Arginine Binding Protein (TmArgBP) as model. Specifically, we mutated TmArgBP Gly52 that assumes a conformation believed to be strictly disallowed for non-Gly residues. Surprisingly, the crystallographic characterization of Gly52Ala TmArgBP indicates that the structural context forces the residue to adopt a non-canonical conformation never observed in any of the high-medium resolution PDB structures. Interestingly, the inspection of this high resolution structure demonstrates that only minor alterations occur. Nevertheless, experiments indicate that Gly52 replacements in TmArgBP produce destabilizations comparable to those observed upon protein truncation or dissection in domains. Notably, we show that force-fields commonly used in computational biology do not reproduce this non-canonical state. Using TmArgBP as model system we here demonstrate that the structural context may force residues to adopt conformations believed to be strictly forbidden and that barely detectable alterations produce major destabilizations. Present findings highlight the role of subtle strains in governing protein stability. A full understanding of these phenomena is essential for an exhaustive comprehension of the factors regulating protein structures.
Organizational Affiliation:
Institute of Biostructures and Bioimaging, CNR, Via Mezzocannone 16, Napoli, Italy.