Affinity maturation, humanization, and co-crystallization of a rabbit anti-human ROR2 monoclonal antibody for therapeutic applications.
Goydel, R.S., Weber, J., Peng, H., Qi, J., Soden, J., Freeth, J., Park, H., Rader, C.(2020) J Biol Chem 295: 5995-6006
- PubMed: 32193207
- DOI: https://doi.org/10.1074/jbc.RA120.012791
- Primary Citation of Related Structures:
6OSH, 6OSN, 6OSV - PubMed Abstract:
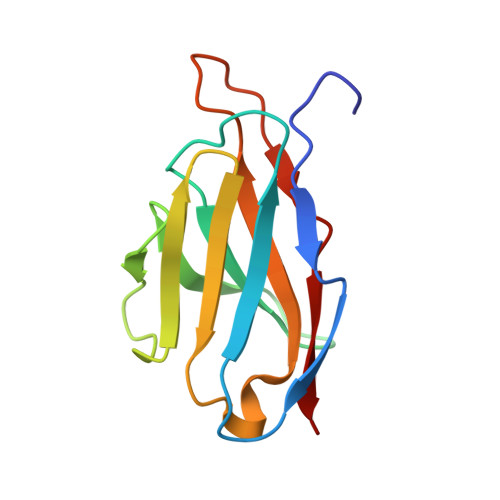
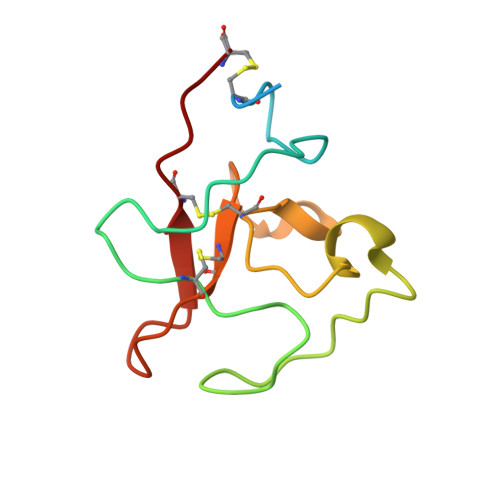
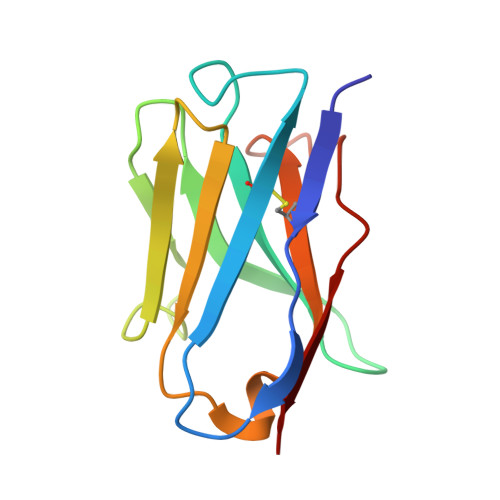
Antibodies are widely used as cancer therapeutics, but their current use is limited by the low number of antigens restricted to cancer cells. A receptor tyrosine kinase, receptor tyrosine kinase-like orphan receptor 2 (ROR2), is normally expressed only during embryogenesis and is tightly down-regulated in postnatal healthy tissues. However, it is up-regulated in a diverse set of hematologic and solid malignancies, thus ROR2 represents a candidate antigen for antibody-based cancer therapy. Here we describe the affinity maturation and humanization of a rabbit mAb that binds human and mouse ROR2 but not human ROR1 or other human cell-surface antigens. Co-crystallization of the parental rabbit mAb in complex with the human ROR2 kringle domain (hROR2-Kr) guided affinity maturation by heavy-chain complementarity-determining region 3 (HCDR3)-focused mutagenesis and selection. The affinity-matured rabbit mAb was then humanized by complementarity-determining region (CDR) grafting and framework fine tuning and again co-crystallized with hROR2-Kr. We show that the affinity-matured and humanized mAb retains strong affinity and specificity to ROR2 and, following conversion to a T cell-engaging bispecific antibody, has potent cytotoxicity toward ROR2-expressing cells. We anticipate that this humanized affinity-matured mAb will find application for antibody-based cancer therapy of ROR2-expressing neoplasms.
Organizational Affiliation:
Department of Immunology and Microbiology, The Scripps Research Institute, Jupiter, Florida 33458.