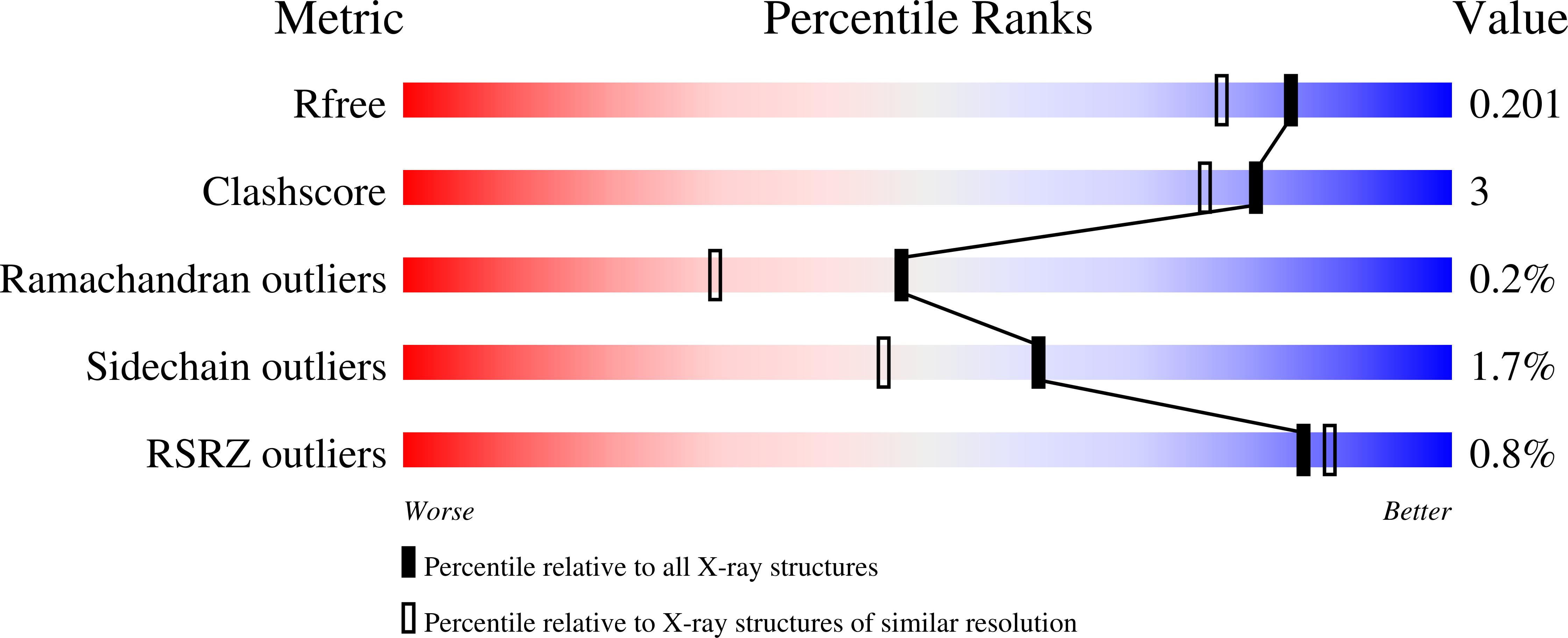
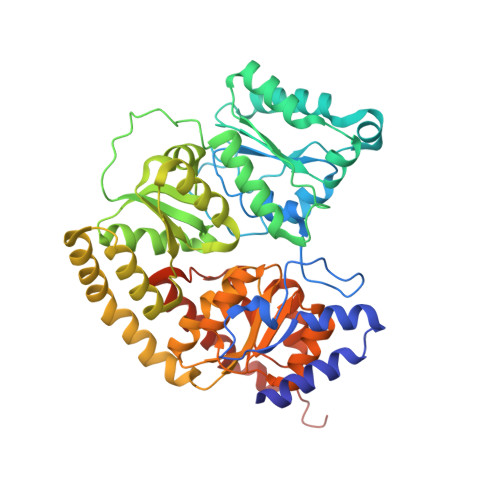
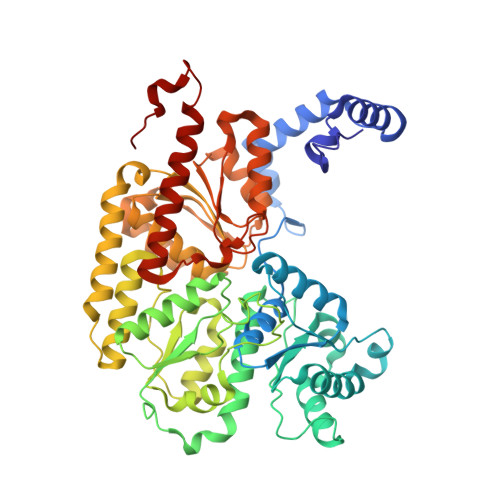
Redox-Dependent Metastability of the Nitrogenase P-Cluster.
Rutledge, H.L., Rittle, J., Williamson, L.M., Xu, W.A., Gagnon, D.M., Tezcan, F.A.(2019) J Am Chem Soc 141: 10091-10098
- PubMed: 31146522
- DOI: https://doi.org/10.1021/jacs.9b04555
- Primary Citation of Related Structures:
6O7L, 6O7M, 6O7N, 6O7O, 6O7P, 6O7Q, 6O7R, 6O7S - PubMed Abstract:
Molybdenum nitrogenase catalyzes the reduction of dinitrogen into ammonia, which requires the coordinated transfer of eight electrons to the active site cofactor (FeMoco) through the intermediacy of an [8Fe-7S] cluster (P-cluster), both housed in the molybdenum-iron protein (MoFeP). Previous studies on MoFeP from two different organisms, Azotobacter vinelandii ( Av) and Gluconacetobacter diazotrophicus ( Gd), have established that the P-cluster is conformationally flexible and can undergo substantial structural changes upon two-electron oxidation to the P OX state, whereby a backbone amidate and an oxygenic residue (Ser or Tyr) ligate to two of the cluster's Fe centers. This redox-dependent change in coordination has been implicated in the conformationally gated electron transfer in nitrogenase. Here, we have investigated the role of the oxygenic ligand in Av MoFeP, which natively contains a Ser ligand (βSer188) to the P-cluster. Three variants were generated in which (1) the oxygenic ligand was eliminated (βSer188Ala), (2) the P-cluster environment was converted to the one in Gd MoFeP (βPhe99Tyr/βSer188Ala), and (3) two oxygenic ligands were simultaneously included (βPhe99Tyr). Our studies have revealed that the P-cluster can become compositionally labile upon oxidation and reversibly lose one or two Fe centers in the absence of the oxygenic ligand, while still retaining wild-type-like dinitrogen reduction activity. Our findings also suggest that Av and Gd MoFePs evolved with specific preferences for Ser and Tyr ligands, respectively, and that the structural control of these ligands must extend beyond the primary and secondary coordination spheres of the P-cluster. The P-cluster adds to the increasing number of examples of inherently labile Fe-S clusters whose compositional instability may be an obligatory feature to enable redox-linked conformational changes to facilitate multielectron redox reactions.
Organizational Affiliation:
Department of Chemistry and Biochemistry , University of California, San Diego , 9500 Gilman Drive , La Jolla , California 92093-0356 , United States.