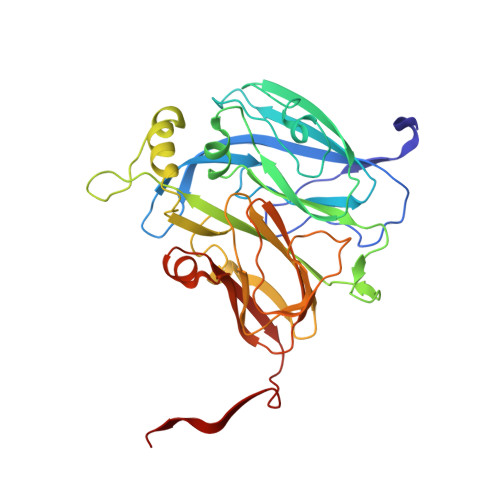
2.85 and 2.99 angstrom resolution structures of 110 kDa nitrite reductase determined by 200 kV cryogenic electron microscopy.
Adachi, N., Yamaguchi, T., Moriya, T., Kawasaki, M., Koiwai, K., Shinoda, A., Yamada, Y., Yumoto, F., Kohzuma, T., Senda, T.(2021) J Struct Biol 213: 107768-107768
- PubMed: 34217801
- DOI: https://doi.org/10.1016/j.jsb.2021.107768
- Primary Citation of Related Structures:
6KNF, 6KNG - PubMed Abstract:
Cu-containing nitrite reductases (NiRs) are 110 kDa enzymes that play central roles in denitrification. Although the NiRs have been well studied, with over 100 Protein Data Bank entries, such issues as crystal packing, photoreduction, and lack of high pH cases have impeded structural analysis of their catalytic mechanisms. Here we show the cryogenic electron microscopy (cryo-EM) structures of Achromobacter cycloclastes NiR (AcNiR) at pH 6.2 and 8.1. The optimization of 3D-reconstruction parameters achieved 2.99 and 2.85 Å resolution. Comprehensive comparisons with cryo-EM and 56 AcNiR crystal structures suggested crystallographic artifacts in residues 185-215 and His255' due to packing and photoreduction, respectively. We used a newly developed map comparison method to detect structural change around the type 2 Cu site. While the theoretical estimation of coordinate errors of cryo-EM structures remains difficult, combined analysis using X-ray and cryo-EM structures will allow deeper insight into the local structural changes of proteins.
Organizational Affiliation:
Structural Biology Research Center, Photon Factory, Institute of Materials Structure Science, High Energy Accelerator Research Organization (KEK), 1-1 Oho, Tsukuba, Ibaraki 305-0801, Japan; Department of Materials Structure Science, School of High Energy Accelerator Science, The Graduate University of Advanced Studies (Soken-dai), 1-1 Oho, Tsukuba, Ibaraki 305-0801, Japan.