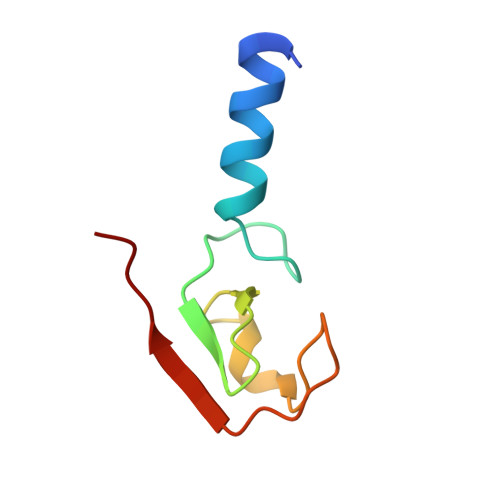
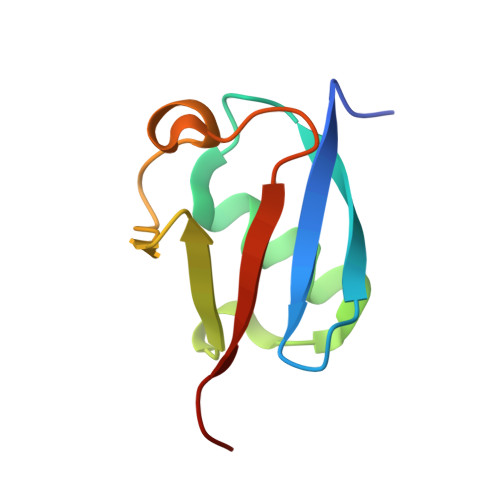
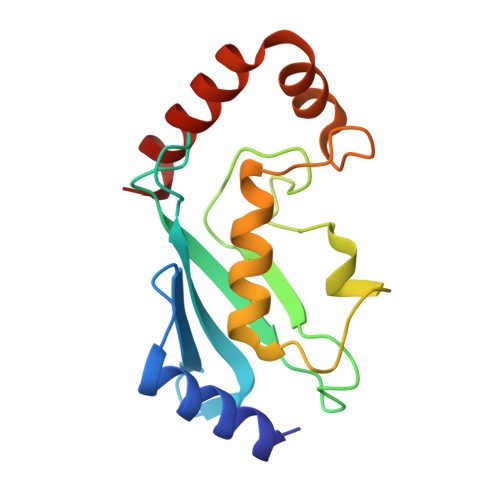
Structural insights into non-covalent ubiquitin activation of the cIAP1-UbcH5B∼ubiquitin complex.
Patel, A., Sibbet, G.J., Huang, D.T.(2019) J Biol Chem 294: 1240-1249
- PubMed: 30523153
- DOI: https://doi.org/10.1074/jbc.RA118.006045
- Primary Citation of Related Structures:
6HPR - PubMed Abstract:
Ubiquitin (Ub)-conjugating enzymes and Ub ligases control protein degradation and regulate many cellular processes in eukaryotes. Cellular inhibitor of apoptosis protein-1 (cIAP1) plays a central role in apoptosis and tumor necrosis factor signaling. It harbors a C-terminal RING domain that homodimerizes to recruit E2∼Ub (where ∼ denotes a thioester bond) complex to catalyze Ub transfer. Noncovalent Ub binding to the backside of the E2 Ub-conjugating enzyme UbcH5 has previously been shown to enhance RING domain activity, but the molecular basis for this enhancement is unclear. To investigate how dimeric cIAP1 RING activates E2∼Ub for Ub transfer and what role noncovalently bound Ub has in Ub transfer, here we determined the crystal structure of the cIAP1 RING dimer bound to both UbcH5B covalently linked to Ub (UbcH5B-Ub) and a noncovalent Ub to 1.7 Å resolution. The structure along with biochemical analyses revealed that the cIAP1 RING domain interacts with UbcH5B-Ub and thereby promotes the formation of a closed UbcH5B-Ub conformation that primes the thioester bond for Ub transfer. We observed that the noncovalent Ub binds to the backside of UbcH5B and abuts UbcH5B's α1β1-loop, which, in turn, stabilizes the closed UbcH5B-Ub conformation. Our results disclose the mechanism by which cIAP1 RING dimer activates UbcH5B∼Ub and indicate that noncovalent Ub binding further stabilizes the cIAP1-UbcH5B∼Ub complex in the active conformation to stimulate Ub transfer.
Organizational Affiliation:
Cancer Research UK Beatson Institute, Garscube Estate, Switchback Road, Glasgow G61 1BD, Scotland, United Kingdom; Institute of Cancer Sciences, University of Glasgow, Glasgow G61 1BD, Scotland, United Kingdom.