hPDE4D2 structure with inhibitor NPD-425
Singh, A.K., Brown, D.G.To be published.
Experimental Data Snapshot
Starting Model: experimental
View more details
Entity ID: 1 | |||||
|---|---|---|---|---|---|
| Molecule | Chains | Sequence Length | Organism | Details | Image |
| cAMP-specific 3',5'-cyclic phosphodiesterase 4D | 364 | Homo sapiens | Mutation(s): 0 Gene Names: PDE4D, DPDE3 EC: 3.1.4.53 | 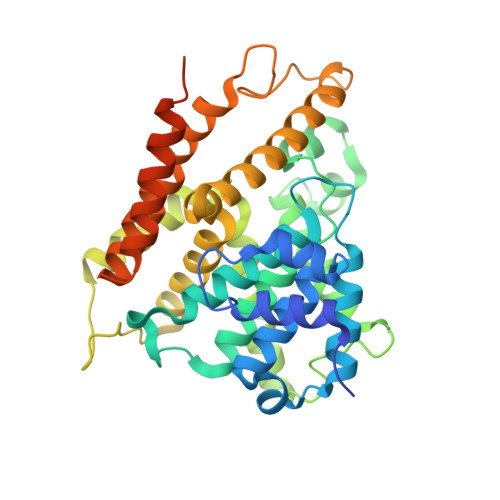 | |
UniProt & NIH Common Fund Data Resources | |||||
Find proteins for Q08499 (Homo sapiens) Explore Q08499 Go to UniProtKB: Q08499 | |||||
PHAROS: Q08499 GTEx: ENSG00000113448 | |||||
Entity Groups | |||||
| Sequence Clusters | 30% Identity50% Identity70% Identity90% Identity95% Identity100% Identity | ||||
| UniProt Group | Q08499 | ||||
Sequence AnnotationsExpand | |||||
| |||||
| Ligands 6 Unique | |||||
|---|---|---|---|---|---|
| ID | Chains | Name / Formula / InChI Key | 2D Diagram | 3D Interactions | |
| E6E (Subject of Investigation/LOI) Query on E6E | JB [auth D], M [auth A], NA [auth C], Z [auth B] | 4-{4-methoxy-3-[(2-methoxyphenyl)methoxy]phenyl}-2-(1-{thieno[3,2-d]pyrimidin-4-yl}piperidin-4-yl)-1,2,4a,5,8,8a-hexahydrophthalazin-1-one C34 H35 N5 O4 S FPUMGIJNOYNQAE-CLJLJLNGSA-N |  | ||
| EPE Query on EPE | BA [auth B], LA [auth C], MB [auth D], N [auth A], Y [auth B] | 4-(2-HYDROXYETHYL)-1-PIPERAZINE ETHANESULFONIC ACID C8 H18 N2 O4 S JKMHFZQWWAIEOD-UHFFFAOYSA-N |  | ||
| PEG Query on PEG | EB [auth D], FB [auth D], KB [auth D] | DI(HYDROXYETHYL)ETHER C4 H10 O3 MTHSVFCYNBDYFN-UHFFFAOYSA-N |  | ||
| ZN Query on ZN | E [auth A], FA [auth C], R [auth B], UA [auth D] | ZINC ION Zn PTFCDOFLOPIGGS-UHFFFAOYSA-N |  | ||
| EDO Query on EDO | AA [auth B] AB [auth D] BB [auth D] CA [auth B] CB [auth D] | 1,2-ETHANEDIOL C2 H6 O2 LYCAIKOWRPUZTN-UHFFFAOYSA-N |  | ||
| MG Query on MG | F [auth A], GA [auth C], S [auth B], VA [auth D] | MAGNESIUM ION Mg JLVVSXFLKOJNIY-UHFFFAOYSA-N |  | ||
| Length ( Å ) | Angle ( ˚ ) |
|---|---|
| a = 98.939 | α = 90 |
| b = 110.812 | β = 90 |
| c = 160.983 | γ = 90 |
| Software Name | Purpose |
|---|---|
| REFMAC | refinement |
| XDS | data reduction |
| autoPROC | data scaling |
| PHASER | phasing |