Bivalent Inhibitor UNC4512 Bound to the TAF1 Bromodomain Tandem
Mathea, S., Suh, J.L., Salah, E., Tallant, C., Siejka, P., Pike, A.C.W., von Delft, F., Arrowsmith, C.H., Edwards, A.M., Bountra, C., James, L.I., Frye, S.V., Knapp, S.To be published.
Experimental Data Snapshot
Starting Model: experimental
View more details
Entity ID: 1 | |||||
|---|---|---|---|---|---|
| Molecule | Chains | Sequence Length | Organism | Details | Image |
| Transcription initiation factor TFIID subunit 1 | A [auth T] | 280 | Homo sapiens | Mutation(s): 0 Gene Names: TAF1, BA2R, CCG1, CCGS, TAF2A EC: 2.3.1.48 (PDB Primary Data), 2.7.11.1 (PDB Primary Data) | 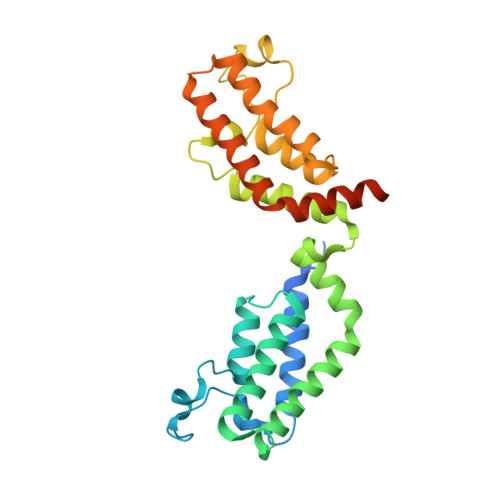 |
UniProt & NIH Common Fund Data Resources | |||||
Find proteins for P21675 (Homo sapiens) Explore P21675 Go to UniProtKB: P21675 | |||||
PHAROS: P21675 GTEx: ENSG00000147133 | |||||
Entity Groups | |||||
| Sequence Clusters | 30% Identity50% Identity70% Identity90% Identity95% Identity100% Identity | ||||
| UniProt Group | P21675 | ||||
Sequence AnnotationsExpand | |||||
| |||||
| Ligands 1 Unique | |||||
|---|---|---|---|---|---|
| ID | Chains | Name / Formula / InChI Key | 2D Diagram | 3D Interactions | |
| DKH (Subject of Investigation/LOI) Query on DKH | B [auth T], C [auth T] | 3-azanyl-2-[3-[2-[2-[2-[2-[2-[2-[2-[2-[2-[2-[2-[2-[2-[2-[2-[2-[2-[[3-azanyl-1-[[2-[[3-methyl-6-[4-methyl-3-(methylsulfonyl-$l^{2}-azanyl)cyclohexa-1,3,5-trien-1-yl]-[1,2,4]triazolo[4,3-b]pyridazin-8-yl]-$l^{2}-azanyl]-2-oxidanylidene-ethyl]amino]-1-oxidanylidene-propan-2-yl]amino]-2-oxidanylidene-ethoxy]ethoxy]ethoxy]ethoxy]ethoxy]ethoxy]ethoxy]ethoxy]ethoxy]ethoxy]ethoxy]ethoxy]ethoxy]ethoxy]ethoxy]ethoxy]ethoxy]propanoylamino]-~{N}-[3-[[3-methyl-6-[4-methyl-3-(methylsulfonylamino)phenyl]-[1,2,4]triazolo[4,3-b]pyridazin-8-yl]amino]-3-oxidanylidene-propyl]propanamide C76 H105 N18 O27 S2 PSQFSMQXCFCIIU-UHFFFAOYSA-N |  | ||
| Length ( Å ) | Angle ( ˚ ) |
|---|---|
| a = 113.573 | α = 90 |
| b = 113.573 | β = 90 |
| c = 72.592 | γ = 120 |
| Software Name | Purpose |
|---|---|
| REFMAC | refinement |
| xia2 | data reduction |
| xia2 | data scaling |
| PHASER | phasing |