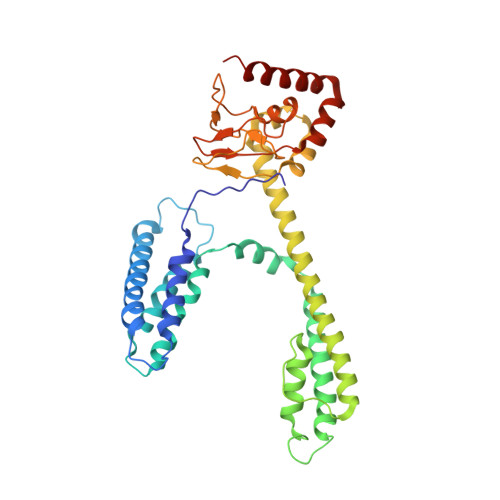
High-Resolution Cryoelectron Microscopy Structure of the Cyclic Nucleotide-Modulated Potassium Channel MloK1 in a Lipid Bilayer.
Kowal, J., Biyani, N., Chami, M., Scherer, S., Rzepiela, A.J., Baumgartner, P., Upadhyay, V., Nimigean, C.M., Stahlberg, H.(2018) Structure 26: 20-27.e3
- PubMed: 29249605
- DOI: https://doi.org/10.1016/j.str.2017.11.012
- Primary Citation of Related Structures:
6EO1 - PubMed Abstract:
Eukaryotic cyclic nucleotide-modulated channels perform their diverse physiological roles by opening and closing their pores to ions in response to cyclic nucleotide binding. We here present a structural model for the cyclic nucleotide-modulated potassium channel homolog from Mesorhizobium loti, MloK1, determined from 2D crystals in the presence of lipids. Even though crystals diffract electrons to only ∼10 Å, using cryoelectron microscopy (cryo-EM) and recently developed computational methods, we have determined a 3D map of full-length MloK1 in the presence of cyclic AMP (cAMP) at ∼4.5 Å isotropic 3D resolution. The structure provides a clear picture of the arrangement of the cyclic nucleotide-binding domains with respect to both the pore and the putative voltage sensor domains when cAMP is bound, and reveals a potential gating mechanism in the context of the lipid-embedded channel.
Organizational Affiliation:
Center for Cellular Imaging and NanoAnalytics (C-CINA), Biozentrum, University of Basel, WRO-1058, Mattenstrasse 26, 4058 Basel, Switzerland.