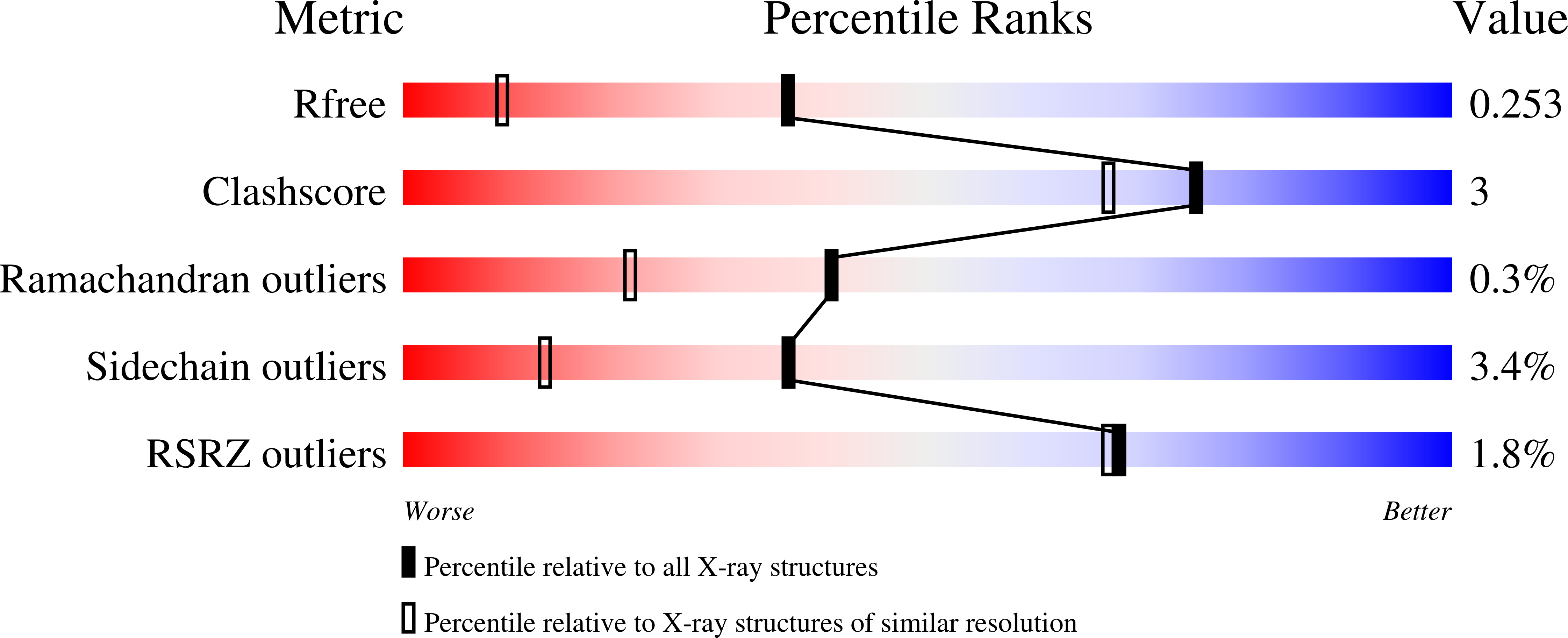
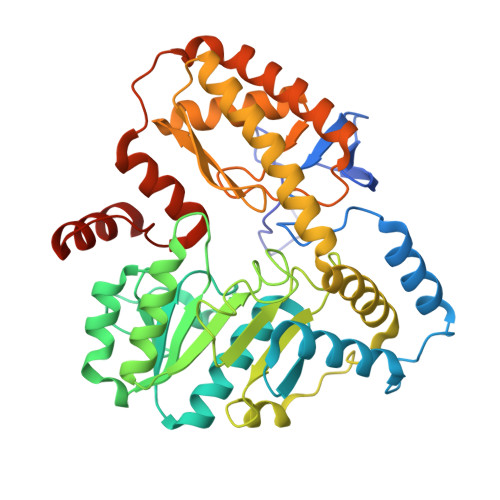
The three-dimensional structure of NeoB: An aminotransferase involved in the biosynthesis of neomycin.
Dow, G.T., Thoden, J.B., Holden, H.M.(2018) Protein Sci 27: 945-956
- PubMed: 29516565
- DOI: https://doi.org/10.1002/pro.3400
- Primary Citation of Related Structures:
6CBK, 6CBL, 6CBM, 6CBN, 6CBO - PubMed Abstract:
The aminoglycoside antibiotics, discovered as natural products in the 1940s, demonstrate a broad antimicrobial spectrum. Due to their nephrotoxic and ototoxic side effects, however, their widespread clinical usage has typically been limited to the treatment of serious infections. Neomycin B, first isolated from strains of Streptomyces in 1948, is one such drug that was approved for human use by the U.S. Food and Drug Administration in 1964. Only within the last 11 years has the biochemical pathway for its production been elaborated, however. Here we present the three-dimensional architecture of NeoB from Streptomyces fradiae, which is a pyridoxal 5'-phosphate or PLP-dependent aminotransferase that functions on two different substrates in neomycin B biosynthesis. For this investigation, four high resolution X-ray structures of NeoB were determined in various complexed states. The overall fold of NeoB is that typically observed for members of the "aspartate aminotransferase" family with the exception of an additional three-stranded antiparallel β-sheet that forms part of the subunit-subunit interface of the dimer. The manner in which the active site of NeoB accommodates quite different substrates has been defined by this investigation. In addition, during the course of this study, we also determined the structure of the aminotransferase GenB1 to high resolution. GenB1 functions as an aminotransferase in gentamicin biosynthesis. Taken together, the structures of NeoB and GenB1, presented here, provide the first detailed descriptions of aminotransferases that specifically function on aldehyde moieties in aminoglycoside biosynthesis.
Organizational Affiliation:
Department of Biochemistry, University of Wisconsin, Madison, WI, 53706.