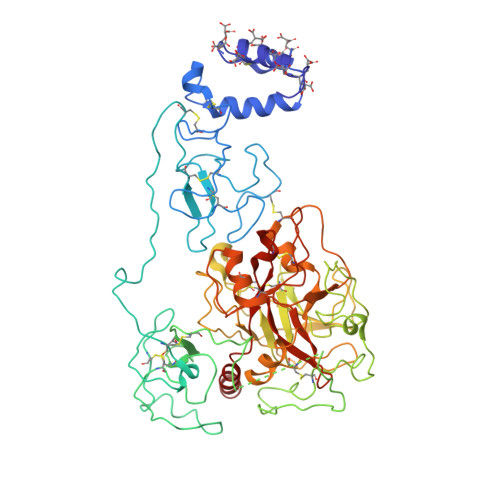
Structure of prothrombin in the closed form reveals new details on the mechanism of activation.
Chinnaraj, M., Chen, Z., Pelc, L.A., Grese, Z., Bystranowska, D., Di Cera, E., Pozzi, N.(2018) Sci Rep 8: 2945-2945
- PubMed: 29440720
- DOI: https://doi.org/10.1038/s41598-018-21304-1
- Primary Citation of Related Structures:
6BJR, 6C2W - PubMed Abstract:
The clotting factor prothrombin exists in equilibrium between closed and open conformations, but the physiological role of these forms remains unclear. As for other allosteric proteins, elucidation of the linkage between molecular transitions and function is facilitated by reagents stabilized in each of the alternative conformations. The open form of prothrombin has been characterized structurally, but little is known about the architecture of the closed form that predominates in solution under physiological conditions. Using X-ray crystallography and single-molecule FRET, we characterize a prothrombin construct locked in the closed conformation through an engineered disulfide bond. The construct: (i) provides structural validation of the intramolecular collapse of kringle-1 onto the protease domain reported recently; (ii) documents the critical role of the linker connecting kringle-1 to kringle-2 in stabilizing the closed form; and (iii) reveals novel mechanisms to shift the equilibrium toward the open conformation. Together with functional studies, our findings define the role of closed and open conformations in the conversion of prothrombin to thrombin and establish a molecular framework for prothrombin activation that rationalizes existing phenotypes associated with prothrombin mutations and points to new strategies for therapeutic intervention.
Organizational Affiliation:
Edward A. Doisy Department of Biochemistry and Molecular Biology, Saint Louis University School of Medicine, St. Louis, MO, 63104, USA.