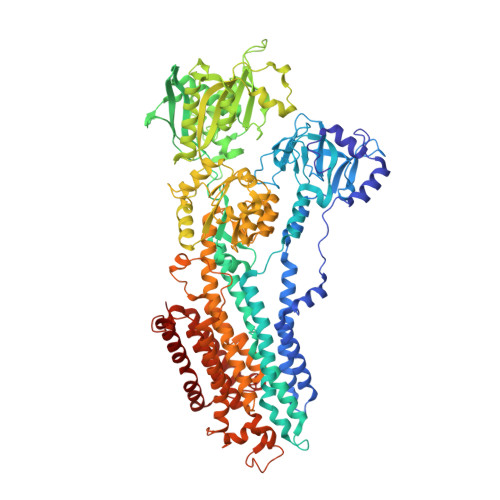
Protein-phospholipid interplay revealed with crystals of a calcium pump.
Norimatsu, Y., Hasegawa, K., Shimizu, N., Toyoshima, C.(2017) Nature 545: 193-198
- PubMed: 28467821
- DOI: https://doi.org/10.1038/nature22357
- Primary Citation of Related Structures:
5XA7, 5XA8, 5XA9, 5XAA, 5XAB - PubMed Abstract:
The lipid bilayer has so far eluded visualization by conventional crystallographic methods, severely limiting our understanding of phospholipid- and protein-phospholipid interactions. Here we describe electron density maps for crystals of Ca 2+ -ATPase in four different states obtained by X-ray solvent contrast modulation. These maps resolve the entire first layer of phospholipids surrounding the transmembrane helices, although less than half of them are hydrogen-bonded to protein residues. Phospholipids follow the movements of associated residues, causing local distortions and changes in thickness of the bilayer. Unexpectedly, the entire protein tilts during the reaction cycle, governed primarily by a belt of Trp residues, to minimize energy costs accompanying the large perpendicular movements of the transmembrane helices. A class of Arg residues extend their side chains through the cytoplasm to exploit phospholipids as anchors for conformational switching. Thus, phospholipid-Arg/Lys and phospholipid-Trp interactions have distinct functional roles in the dynamics of ion pumps and, presumably, membrane proteins in general.
Organizational Affiliation:
Institute of Molecular and Cellular Biosciences, The University of Tokyo, Tokyo 113-0032, Japan.