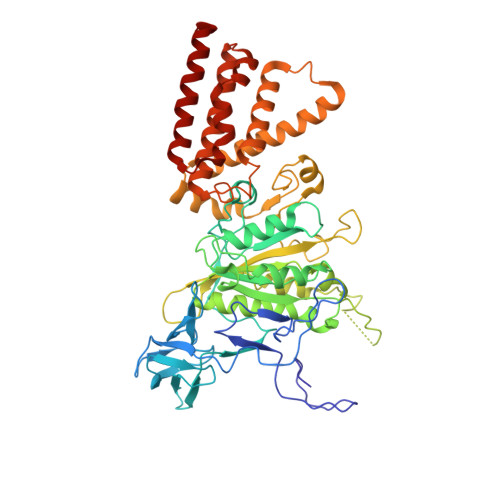
Crystallographic and enzymatic insights into the mechanisms of Mg-ADP inhibition in the A1 complex of the A1AO ATP synthase
Singh, D., Gruber, G.(2018) J Struct Biol 201: 26-35
- PubMed: 29074108
- DOI: https://doi.org/10.1016/j.jsb.2017.10.008
- Primary Citation of Related Structures:
5X09 - PubMed Abstract:
F-ATP synthases are described to have mechanisms which regulate the unnecessary depletion of ATP pool during an energy limited state of the cell. Mg-ADP inhibition is one of the regulatory features where Mg-ADP gets entrapped in the catalytic site, preventing the binding of ATP and further inhibiting ATP hydrolysis. Knowledge about the existence and regulation of the related archaeal-type A 1 A O ATP synthases (A 3 B 3 CDE 2 FG 2 ac) is limited. We demonstrate MgADP inhibition of the enzymatically active A 3 B 3 D- and A 3 B 3 DF complexes of Methanosarcina mazei Gö1 A-ATP synthase and reveal the importance of the amino acids P235 and S238 inside the P-loop (GPFGSGKTV) of the catalytic A subunit. Substituting these two residues by the respective P-loop residues alanine and cysteine (GAFGCGKTV) of the related eukaryotic V-ATPase increases significantly the ATPase activity of the enzyme variant and abolishes MgADP inhibition. The atomic structure of the P235A, S238C double mutant of subunit A of the Pyrococcus horikoshii OT3 A-ATP synthase provides details of how these critical residues affect nucleotide-binding and ATP hydrolysis in this molecular engine. The qualitative data are confirmed by quantitative results derived from fluorescence correlation spectroscopy experiments.
Organizational Affiliation:
Nanyang Technological University, School of Biological Sciences, 60 Nanyang Drive, Singapore 637551, Republic of Singapore.