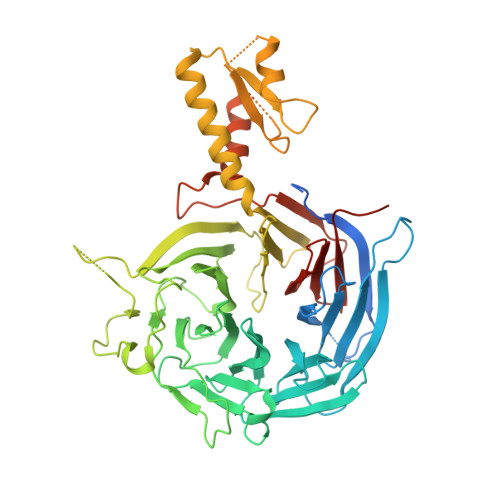
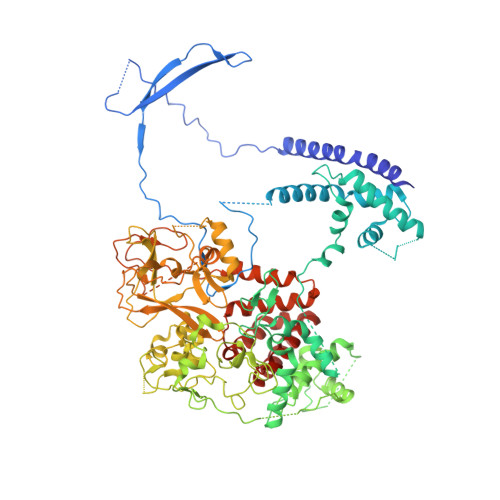
An Evolutionarily Conserved Structural Platform for PRC2 Inhibition by a Class of Ezh2 Inhibitors.
Bratkowski, M., Yang, X., Liu, X.(2018) Sci Rep 8: 9092-9092
- PubMed: 29904056
- DOI: https://doi.org/10.1038/s41598-018-27175-w
- Primary Citation of Related Structures:
5WF7, 5WFC, 5WFD, 5WG6 - PubMed Abstract:
Polycomb repressive complex 2 (PRC2) mediates trimethylation of histone H3K27 (H3K27me3), an epigenetic hallmark for repressed chromatin. Overactive mutants of the histone lysine methyltransferase subunit of PRC2, Ezh2, are found in various types of cancers. Pyridone-containing inhibitors such as GSK126 compete with S-adenosylmethionine (SAM) for Ezh2 binding and effectively inhibit PRC2 activity. PRC2 from the thermophilic fungus Chaetomium thermophilum (ct) is functionally similar to the human version in several regards and has the added advantage of producing high-resolution crystal structures, although inhibitor-bound structures of human or human/chameleon PRC2 are also available at up to 2.6 Å resolution. We solved crystal structures of both human and ctPRC2 bound to GSK126 and the structurally similar inhibitor GSK343. While the two organisms feature a disparate degree of inhibitor potency, surprisingly, GSK126 binds in a similar manner in both structures. Structure-guided protein engineering of the drug binding pocket allowed us to introduce humanizing mutations into ctEzh2 to produce a ctPRC2 variant that is more susceptible to GSK126 inhibition. Additional analysis indicated that an evolutionarily conserved structural platform dictates a unique mode of GSK126 binding, suggesting a mechanism of drug selectivity. The existing drug scaffold may thus be used to probe the function and cellular regulation of PRC2 in a wide spectrum of organisms, ranging from fungi to humans.
Organizational Affiliation:
Cecil H. and Ida Green Center for Reproductive Biology Sciences and Division of Basic Research, Department of Obstetrics and Gynecology, Department of Biophysics, UT Southwestern Medical Center, Dallas, TX, 75390, USA.