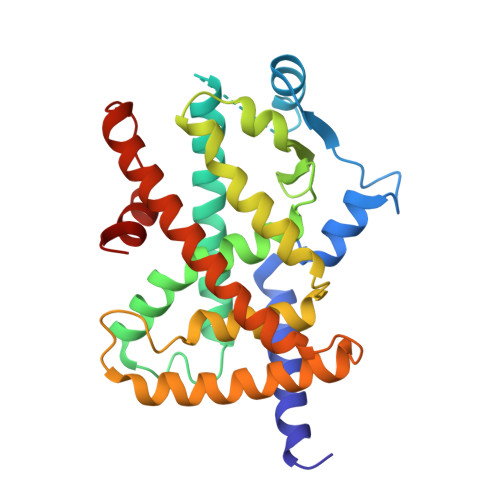
X-ray crystal structure of rivoglitazone bound to PPAR gamma and PPAR subtype selectivity of TZDs.
Rajapaksha, H., Bhatia, H., Wegener, K., Petrovsky, N., Bruning, J.B.(2017) Biochim Biophys Acta 1861: 1981-1991
- PubMed: 28499821
- DOI: https://doi.org/10.1016/j.bbagen.2017.05.008
- Primary Citation of Related Structures:
5U5L - PubMed Abstract:
Thiazolidinedione (TZD) compounds targeting the nuclear receptor peroxisome proliferator-activated receptor gamma (PPARγ) demonstrate unique benefits for the treatment of insulin resistance and type II diabetes. TZDs include rosiglitazone, pioglitazone and rivoglitazone, with the latter being the most potent. The TZDs are only marginally selective for the therapeutic target PPARγ as they also activate PPARα and PPARδ homologues to varying degrees, causing off-target effects. While crystal structures for TZD compounds in complex with PPARγ are available, minimal structural information is available for TZDs bound to PPARα and PPARδ. This paucity of structural information has hampered the determination of precise structural mechanisms involved in TZD selectivity between PPARs. To help address these questions molecular dynamic simulations were performed of rosiglitazone, pioglitazone and rivoglitazone in complex with PPARα, PPARδ, and PPARγ in order to better understand the mechanisms of PPAR selectivity. The simulations revealed that TZD interactions with residues Tyr314 and Phe318 of PPARα and residues Phe291 and Thr253 of PPARδ as well as the omega loop, are key determinants of TZD receptor selectivity. Notably, in this study, we solve the first X-ray crystal structure of rivoglitazone bound to any PPAR. Rivoglitazone forms a unique hydrogen bond network with the residues of the PPARγ co-activator binding surface (known as AF2) and makes more extensive contacts with helix 3 and the β-sheet as compared to model TZD compounds such as rosiglitazone.
Organizational Affiliation:
Department of Diabetes and Endocrinology, School of Medicine, Flinders University, Adelaide, South Australia 5042, Australia.