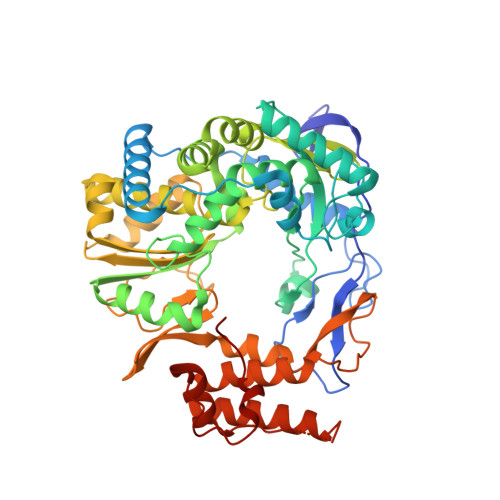
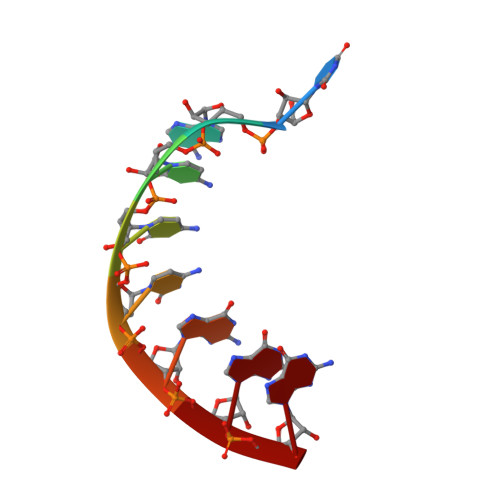
Norovirus RNA-dependent RNA polymerase: A computational study of metal-binding preferences.
Shaik, M.M., Bhattacharjee, N., Feliks, M., Ng, K.K., Field, M.J.(2017) Proteins 85: 1435-1445
- PubMed: 28383118
- DOI: https://doi.org/10.1002/prot.25304
- Primary Citation of Related Structures:
5TSN - PubMed Abstract:
Norovirus (NV) RNA-dependent RNA polymerase (RdRP) is essential for replicating the genome of the virus, which makes this enzyme a key target for the development of antiviral agents against NV gastroenteritis. In this work, a complex of NV RdRP bound to manganese ions and an RNA primer-template duplex was investigated using X-ray crystallography and hybrid quantum chemical/molecular mechanical simulations. Experimentally, the complex crystallized in a tetragonal crystal form. The nature of the primer/template duplex binding in the resulting structure indicates that the complex is a closed back-tracked state of the enzyme, in which the 3'-end of the primer occupies the position expected for the post-incorporated nucleotide before translocation. Computationally, it is found that the complex can accept a range of divalent metal cations without marked distortions in the active site structure. The highest binding energy is for copper, followed closely by manganese and iron, and then by zinc, nickel, and cobalt. Proteins 2017; 85:1435-1445. © 2017 Wiley Periodicals, Inc.
Organizational Affiliation:
Division of Molecular Medicine, Boston Children's Hospital, Boston, Massachusetts, 02115.