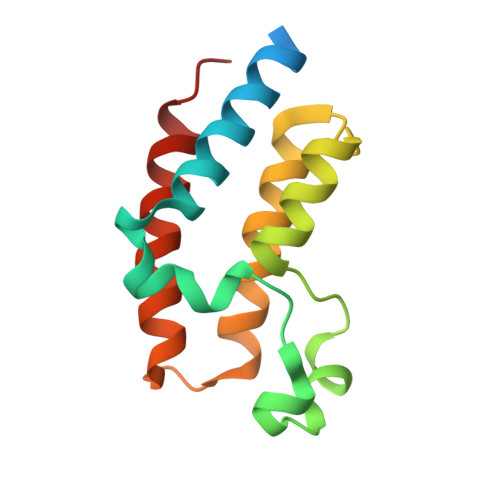
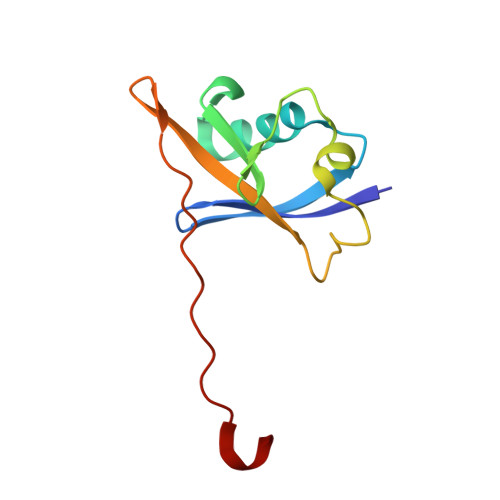
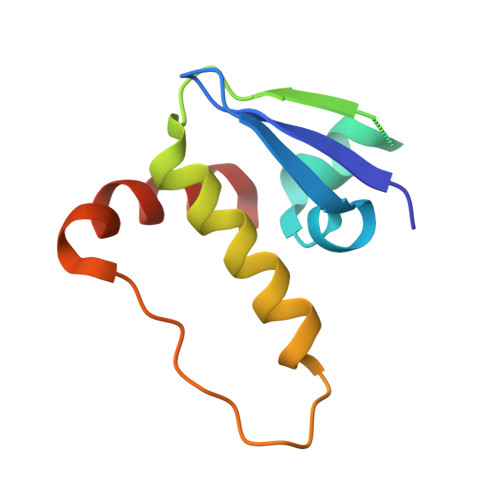
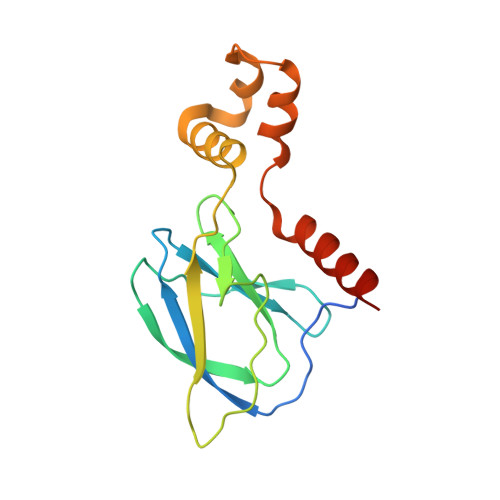
Structural basis of PROTAC cooperative recognition for selective protein degradation.
Gadd, M.S., Testa, A., Lucas, X., Chan, K.H., Chen, W., Lamont, D.J., Zengerle, M., Ciulli, A.(2017) Nat Chem Biol 13: 514-521
- PubMed: 28288108
- DOI: https://doi.org/10.1038/nchembio.2329
- Primary Citation of Related Structures:
5T35 - PubMed Abstract:
Inducing macromolecular interactions with small molecules to activate cellular signaling is a challenging goal. PROTACs (proteolysis-targeting chimeras) are bifunctional molecules that recruit a target protein in proximity to an E3 ubiquitin ligase to trigger protein degradation. Structural elucidation of the key ternary ligase-PROTAC-target species and its impact on target degradation selectivity remain elusive. We solved the crystal structure of Brd4 degrader MZ1 in complex with human VHL and the Brd4 bromodomain (Brd4 BD2 ). The ligand folds into itself to allow formation of specific intermolecular interactions in the ternary complex. Isothermal titration calorimetry studies, supported by surface mutagenesis and proximity assays, are consistent with pronounced cooperative formation of ternary complexes with Brd4 BD2 . Structure-based-designed compound AT1 exhibits highly selective depletion of Brd4 in cells. Our results elucidate how PROTAC-induced de novo contacts dictate preferential recruitment of a target protein into a stable and cooperative complex with an E3 ligase for selective degradation.
Organizational Affiliation:
Division of Biological Chemistry and Drug Discovery, School of Life Sciences, University of Dundee, Dow Street, Dundee, DD1 5EH, Scotland, UK.