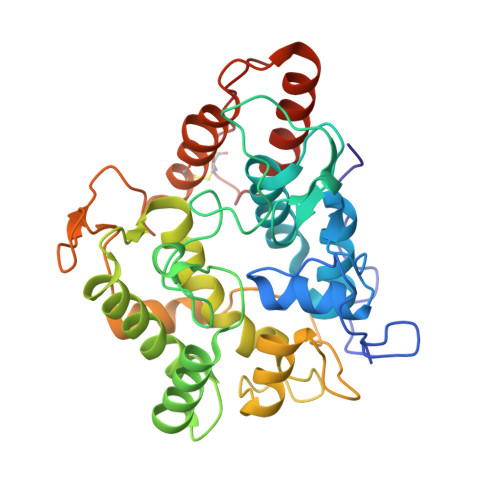
Structural Insights into the Substrate Promiscuity of a Laboratory-Evolved Peroxygenase.
Ramirez-Escudero, M., Molina-Espeja, P., Gomez de Santos, P., Hofrichter, M., Sanz-Aparicio, J., Alcalde, M.(2018) ACS Chem Biol 13: 3259-3268
- PubMed: 30376293
- DOI: https://doi.org/10.1021/acschembio.8b00500
- Primary Citation of Related Structures:
5OXT, 5OXU, 5OY1, 5OY2, 6EKW, 6EKX, 6EKY, 6EKZ, 6EL0, 6EL4 - PubMed Abstract:
Because of their minimal requirements, substrate promiscuity and product selectivity, fungal peroxygenases are now considered to be the jewel in the crown of C-H oxyfunctionalization biocatalysts. In this work, the crystal structure of the first laboratory-evolved peroxygenase expressed by yeast was determined at a resolution of 1.5 Å. Notable differences were detected between the evolved and native peroxygenase from Agrocybe aegerita, including the presence of a full N-terminus and a broader heme access channel due to the mutations that accumulated through directed evolution. Further mutagenesis and soaking experiments with a palette of peroxygenative and peroxidative substrates suggested dynamic trafficking through the heme channel as the main driving force for the exceptional substrate promiscuity of peroxygenase. Accordingly, this study provides the first structural evidence at an atomic level regarding the mode of substrate binding for this versatile biocatalyst, which is discussed within a biological and chemical context.
Organizational Affiliation:
Department of Crystallography & Structural Biology, Institute of Physical Chemistry "Rocasolano" , CSIC , 28006 Madrid , Spain.