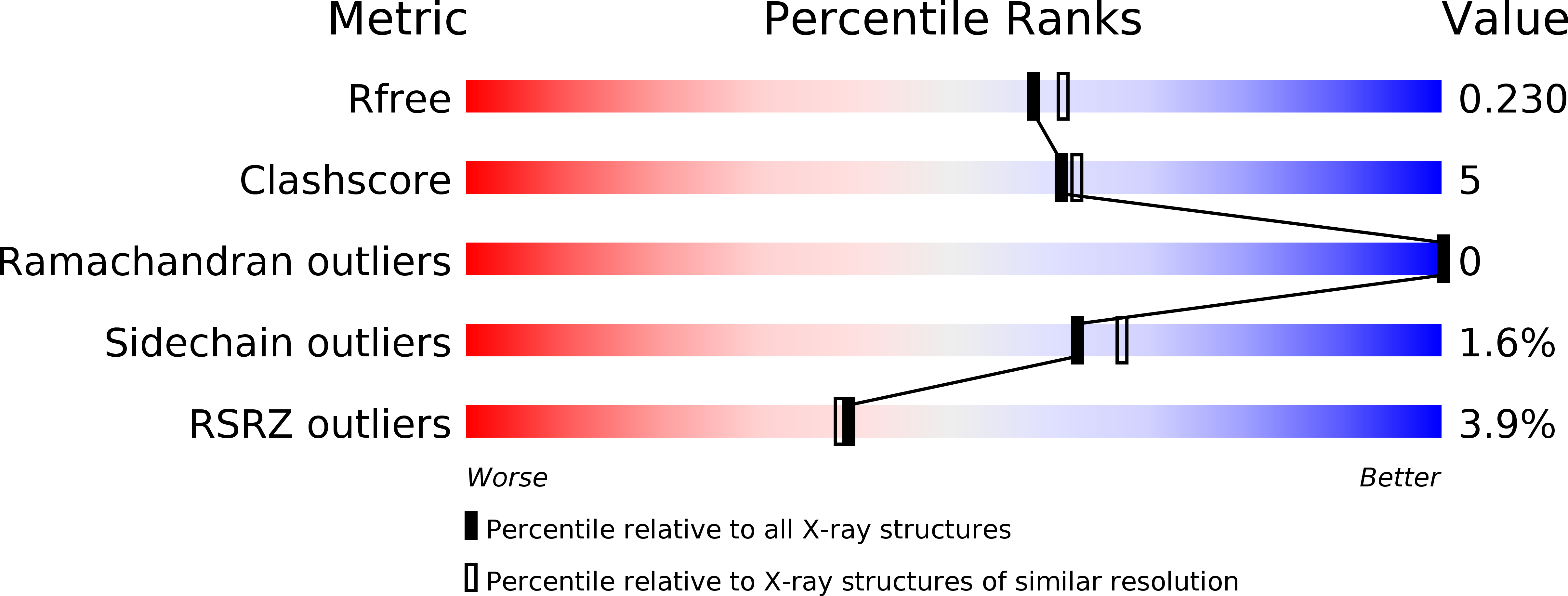
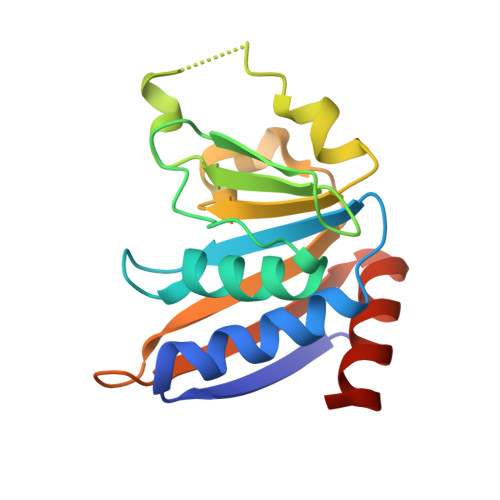
Expanding the Kinome World: A New Protein Kinase Family Widely Conserved in Bacteria.
Nguyen, H.A., El Khoury, T., Guiral, S., Laaberki, M.H., Candusso, M.P., Galisson, F., Foucher, A.E., Kesraoui, S., Ballut, L., Vallet, S., Orelle, C., Zucchini, L., Martin, J., Page, A., Attieh, J., Aghajari, N., Grangeasse, C., Jault, J.M.(2017) J Mol Biol 429: 3056-3074
- PubMed: 28890133
- DOI: https://doi.org/10.1016/j.jmb.2017.08.016
- Primary Citation of Related Structures:
5MVR, 5NP9 - PubMed Abstract:
Fine tuning of signaling pathways is essential for cells to cope with sudden environmental variations. This delicate balance is maintained in particular by protein kinases that control the activity of target proteins by reversible phosphorylation. In addition to homologous eukaryotic enzymes, bacteria have evolved some specific Ser/Thr/Tyr protein kinases without any structural resemblance to their eukaryotic counterparts. Here, we show that a previously identified family of ATPases, broadly conserved among bacteria, is in fact a new family of protein kinases with a Ser/Thr/Tyr kinase activity. A prototypic member of this family, YdiB from Bacillus subtilis, is able to autophosphorylate and to phosphorylate a surrogate substrate, the myelin basic protein. Two crystal structures of YdiB were solved (1.8 and 2.0Å) that display a unique ATP-binding fold unrelated to known protein kinases, although a conserved HxD motif is reminiscent of that found in Hanks-type protein kinases. The effect of mutations of conserved residues further highlights the unique nature of this new protein kinase family that we name ubiquitous bacterial kinase. We investigated the cellular role of YdiB and showed that a ∆ydiB mutant was more sensitive to paraquat treatment than the wild type, with ~13% of cells with an aberrant morphology. In addition, YdiE, which is known to participate with both YdiC and YdiB in an essential chemical modification of some specific tRNAs, is phosphorylated in vitro by YdiB. These results expand the boundaries of the bacterial kinome and support the involvement of YdiB in protein translation and resistance to oxidative stress in B. subtilis.
Organizational Affiliation:
Institut de Biologie Structurale, Université Joseph Fourier Grenoble 1, UMR5075 CNRS/CEA/UJF, 41 rue Jules Horowitz, 38027 Grenoble Cedex 1, France.