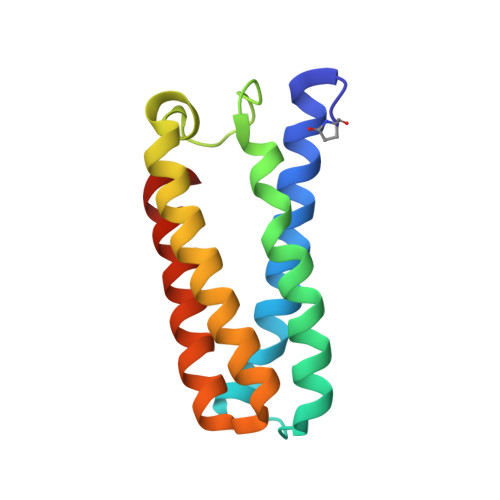
Distinguishing Nitro vs Nitrito Coordination in Cytochrome c' Using Vibrational Spectroscopy and Density Functional Theory.
Nilsson, Z.N., Mandella, B.L., Sen, K., Kekilli, D., Hough, M.A., Moenne-Loccoz, P., Strange, R.W., Andrew, C.R.(2017) Inorg Chem 56: 13205-13213
- PubMed: 29053273
- DOI: https://doi.org/10.1021/acs.inorgchem.7b01945
- Primary Citation of Related Structures:
5NC0, 5NGX - PubMed Abstract:
Nitrite coordination to heme cofactors is a key step in the anaerobic production of the signaling molecule nitric oxide (NO). An ambidentate ligand, nitrite has the potential to coordinate via the N- (nitro) or O- (nitrito) atoms in a manner that can direct its reactivity. Distinguishing nitro vs nitrito coordination, along with the influence of the surrounding protein, is therefore of particular interest. In this study, we probed Fe(III) heme-nitrite coordination in Alcaligenes xylosoxidans cytochrome c' (AXCP), an NO carrier that excludes anions in its native state but that readily binds nitrite (K d ∼ 0.5 mM) following a distal Leu16 → Gly mutation to remove distal steric constraints. Room-temperature resonance Raman spectra (407 nm excitation) identify ν(Fe-NO 2 ), δ(ONO), and ν s (NO 2 ) nitrite ligand vibrations in solution. Illumination with 351 nm UV light results in photoconversion to {FeNO} 6 and {FeNO} 7 states, enabling FTIR measurements to distinguish ν s (NO 2 ) and ν as (NO 2 ) vibrations from differential spectra. Density functional theory calculations highlight the connections between heme environment, nitrite coordination mode, and vibrational properties and confirm that nitrite binds to L16G AXCP exclusively through the N atom. Efforts to obtain the nitrite complex crystal structure were hampered by photochemistry in the X-ray beam. Although low dose crystal structures could be modeled with a mixed nitrite (nitro)/H 2 O distal population, their photosensitivity and partial occupancy underscores the value of the vibrational approach. Overall, this study sheds light on steric determinants of heme-nitrite binding and provides vibrational benchmarks for future studies of heme protein nitrite reactions.
Organizational Affiliation:
Department of Chemistry and Biochemistry, Eastern Oregon University , La Grande, Oregon 97850, United States.