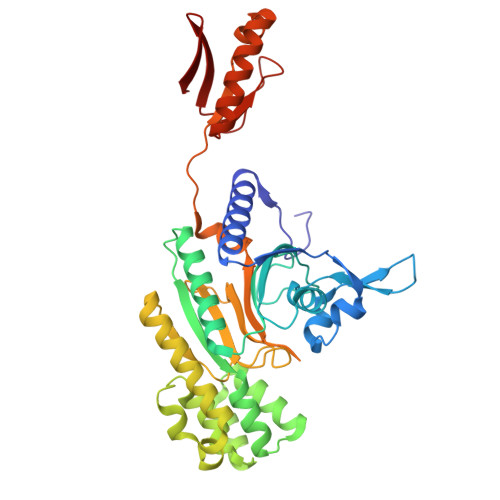
Kinetics and Structure of a Cold-Adapted Hetero-Octameric ATP Phosphoribosyltransferase.
Stroek, R., Ge, Y., Talbot, P.D., Glok, M.K., Bernas, K.E., Thomson, C.M., Gould, E.R., Alphey, M.S., Liu, H., Florence, G.J., Naismith, J.H., da Silva, R.G.(2017) Biochemistry 56: 793-803
- PubMed: 28092443
- DOI: https://doi.org/10.1021/acs.biochem.6b01138
- Primary Citation of Related Structures:
5M8H - PubMed Abstract:
Adenosine 5'-triphosphate phosphoribosyltransferase (ATPPRT) catalyzes the first step in histidine biosynthesis, the condensation of ATP and 5-phospho-α-d-ribosyl-1-pyrophosphate to generate N 1 -(5-phospho-β-d-ribosyl)-ATP and inorganic pyrophosphate. The enzyme is allosterically inhibited by histidine. Two forms of ATPPRT, encoded by the hisG gene, exist in nature, depending on the species. The long form, HisG L , is a single polypeptide chain with catalytic and regulatory domains. The short form, HisG S , lacks a regulatory domain and cannot bind histidine. HisG S instead is found in complex with a regulatory protein, HisZ, constituting the ATPPRT holoenzyme. HisZ triggers HisG S catalytic activity while rendering it sensitive to allosteric inhibition by histidine. Until recently, HisG S was thought to be catalytically inactive without HisZ. Here, recombinant HisG S and HisZ from the psychrophilic bacterium Psychrobacter arcticus were independently overexpressed and purified. The crystal structure of P. arcticus ATPPRT was determined at 2.34 Å resolution, revealing an equimolar HisG S -HisZ hetero-octamer. Steady-state kinetics indicate that both the ATPPRT holoenzyme and HisG S are catalytically active. Surprisingly, HisZ confers only a modest 2-4-fold increase in k cat . Reaction profiles for both enzymes cannot be distinguished by 31 P nuclear magnetic resonance, indicating that the same reaction is catalyzed. The temperature dependence of k cat shows deviation from Arrhenius behavior at 308 K with the holoenzyme. Interestingly, such deviation is detected only at 313 K with HisG S . Thermal denaturation by CD spectroscopy resulted in T m 's of 312 and 316 K for HisZ and HisG S , respectively, suggesting that HisZ renders the ATPPRT complex more thermolabile. This is the first characterization of a psychrophilic ATPPRT.
Organizational Affiliation:
School of Engineering and Applied Science, Rotterdam University of Applied Science , G. J. de Jonghweg 4-6, 3015 GG Rotterdam, The Netherlands.