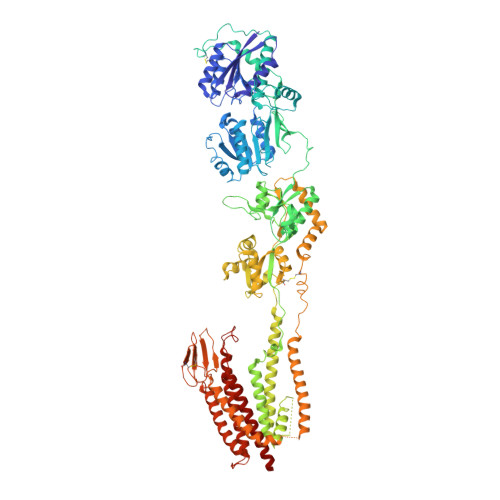
Elucidation of AMPA receptor-stargazin complexes by cryo-electron microscopy.
Twomey, E.C., Yelshanskaya, M.V., Grassucci, R.A., Frank, J., Sobolevsky, A.I.(2016) Science 353: 83-86
- PubMed: 27365450
- DOI: https://doi.org/10.1126/science.aaf8411
- Primary Citation of Related Structures:
5KBS, 5KBT, 5KBU, 5KBV - PubMed Abstract:
AMPA-subtype ionotropic glutamate receptors (AMPARs) mediate fast excitatory neurotransmission and contribute to high cognitive processes such as learning and memory. In the brain, AMPAR trafficking, gating, and pharmacology is tightly controlled by transmembrane AMPAR regulatory proteins (TARPs). Here, we used cryo-electron microscopy to elucidate the structural basis of AMPAR regulation by one of these auxiliary proteins, TARP γ2, or stargazin (STZ). Our structures illuminate the variable interaction stoichiometry of the AMPAR-TARP complex, with one or two TARP molecules binding one tetrameric AMPAR. Analysis of the AMPAR-STZ binding interfaces suggests that electrostatic interactions between the extracellular domains of AMPAR and STZ play an important role in modulating AMPAR function through contact surfaces that are conserved across AMPARs and TARPs. We propose a model explaining how TARPs stabilize the activated state of AMPARs and how the interactions between AMPARs and their auxiliary proteins control fast excitatory synaptic transmission.
Organizational Affiliation:
Department of Biochemistry and Molecular Biophysics, Columbia University, 650 West 168th Street, New York, NY 10032, USA. Integrated Program in Cellular, Molecular, and Biomedical Studies, Columbia University, 650 West 168th Street, New York, NY 10032, USA.