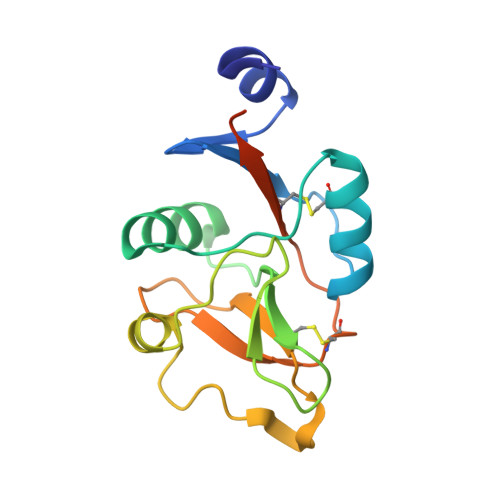
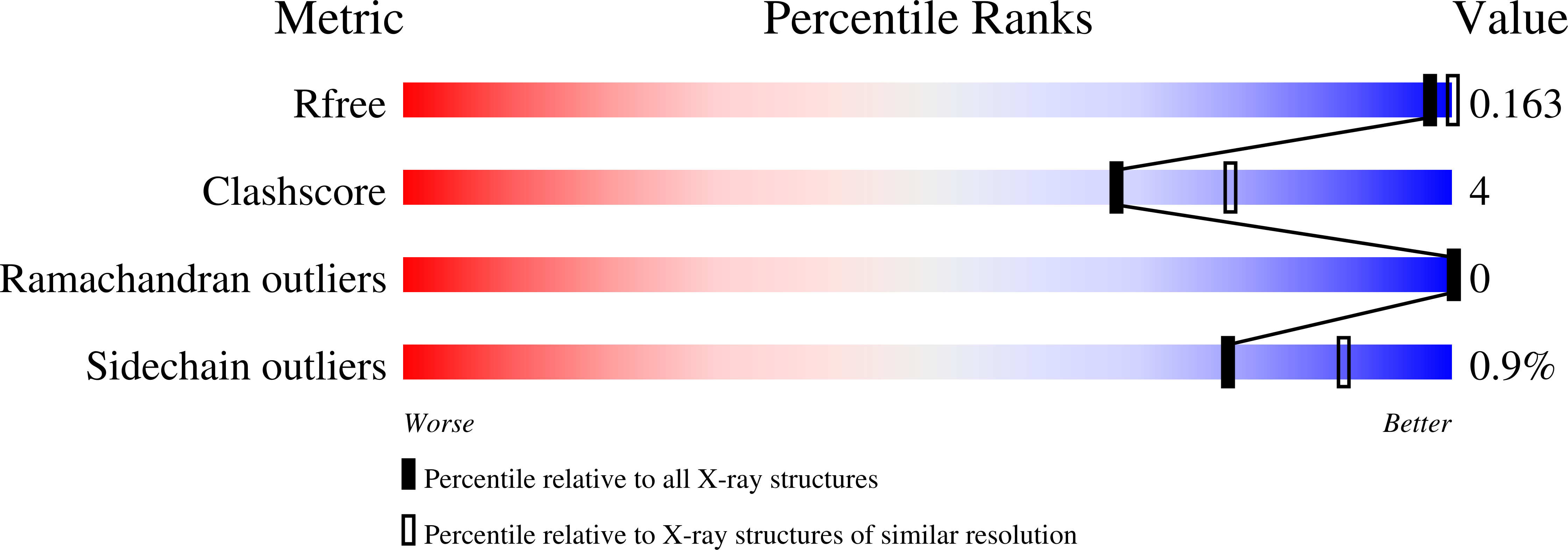Bacterial Polysaccharide Specificity of the Pattern Recognition Receptor Langerin Is Highly Species-dependent.
Hanske, J., Schulze, J., Aretz, J., McBride, R., Loll, B., Schmidt, H., Knirel, Y., Rabsch, W., Wahl, M.C., Paulson, J.C., Rademacher, C.(2017) J Biol Chem 292: 862-871
- PubMed: 27903635
- DOI: https://doi.org/10.1074/jbc.M116.751750
- Primary Citation of Related Structures:
5K8Y, 5M62 - PubMed Abstract:
The recognition of pathogen surface polysaccharides by glycan-binding proteins is a cornerstone of innate host defense. Many members of the C-type lectin receptor family serve as pattern recognition receptors facilitating pathogen uptake, antigen processing, and immunomodulation. Despite the high evolutionary pressure in host-pathogen interactions, it is still widely assumed that genetic homology conveys similar specificities. Here, we investigate the ligand specificities of the human and murine forms of the myeloid C-type lectin receptor langerin for simple and complex ligands augmented by structural insight into murine langerin. Although the two homologs share the same three-dimensional structure and recognize simple ligands identically, a screening of more than 300 bacterial polysaccharides revealed highly diverging avidity and selectivity for larger and more complex glycans. Structural and evolutionary conservation analysis identified a highly variable surface adjacent to the canonic binding site, potentially forming a secondary site of interaction for large glycans.
Organizational Affiliation:
From the Department of Biomolecular Systems, Max Planck Institute of Colloids and Interfaces, Potsdam 14424, Germany.