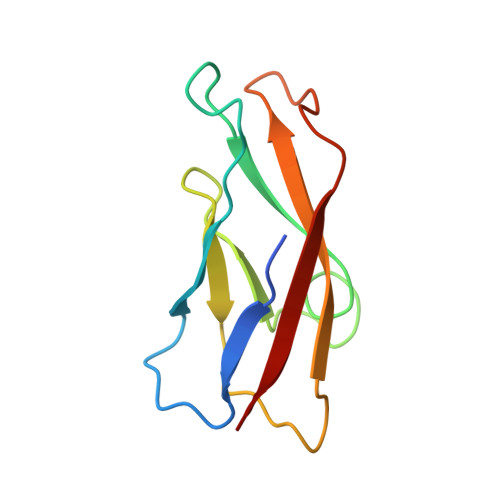
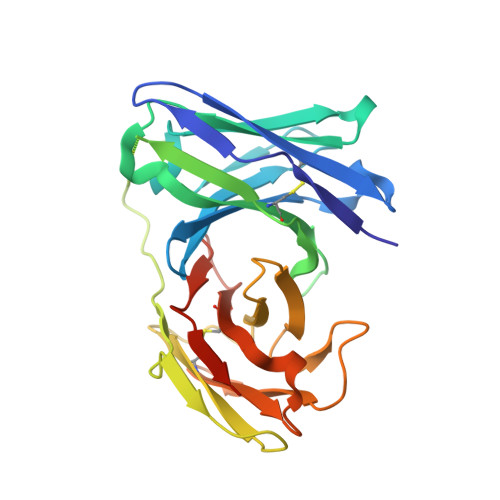
Disruption of cell adhesion by an antibody targeting the cell-adhesive intermediate (X-dimer) of human P-cadherin
Kudo, S., Caaveiro, J.M.M., Nagatoishi, S., Miyafusa, T., Matsuura, T., Sudou, Y., Tsumoto, K.(2017) Sci Rep 7: 39518-39518
- PubMed: 28045038
- DOI: https://doi.org/10.1038/srep39518
- Primary Citation of Related Structures:
5JYL, 5JYM - PubMed Abstract:
Human P-cadherin is a cell adhesion protein of the family of classical cadherins, the overexpression of which is correlated with poor prognosis in various types of cancer. Antibodies inhibiting cell-cell adhesion mediated by P-cadherin show clear therapeutic effect, although the mechanistic basis explaining their effectiveness is still unclear. Based on structural, physicochemical, and functional analyses, we have elucidated the molecular mechanism of disruption of cell adhesion by antibodies targeting human P-cadherin. Herein we have studied three different antibodies, TSP5, TSP7, and TSP11, each recognizing a different epitope on the surface of the cell-adhesive domain (EC1). Although all these three antibodies recognized human P-cadherin with high affinity, only TSP7 disrupted cell adhesion. Notably, we demonstrated that TSP7 abolishes cell adhesion by disabling the so-called X-dimer (a kinetic adhesive intermediate), in addition to disrupting the strand-swap dimer (the final thermodynamic state). The inhibition of the X-dimer was crucial for the overall inhibitory effect, raising the therapeutic value of a kinetic intermediary not only for preventing, but also for reversing, cell adhesion mediated by a member of the classical cadherin family. These findings should help to design more innovative and effective therapeutic solutions targeting human P-cadherin.
Organizational Affiliation:
Department of Chemistry &Biotechnology, School of Engineering, The University of Tokyo, Tokyo 108-8639, Japan.